See the script ./sandbox/coupling-graphs/coupling-graphs-1.R for the implementation of the graphs.
Graph 1
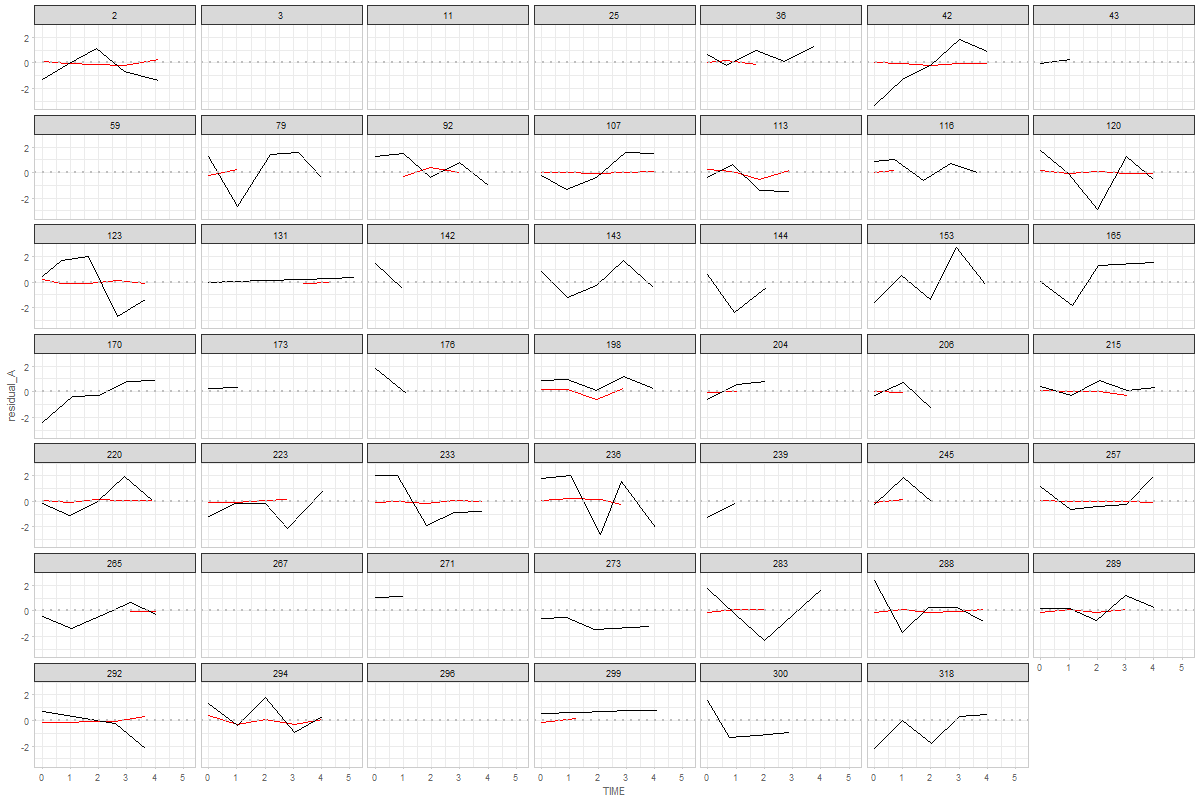 The graph show 48 random individuals (males) from MAP study. Red indicates process A (physical), black represents process B (cognitive). If a person has less then two data points no lines are drawn.
The graph show 48 random individuals (males) from MAP study. Red indicates process A (physical), black represents process B (cognitive). If a person has less then two data points no lines are drawn.
on more similar scale than original metric
not really sure what this means. Could you please elaborate?
Graph 2
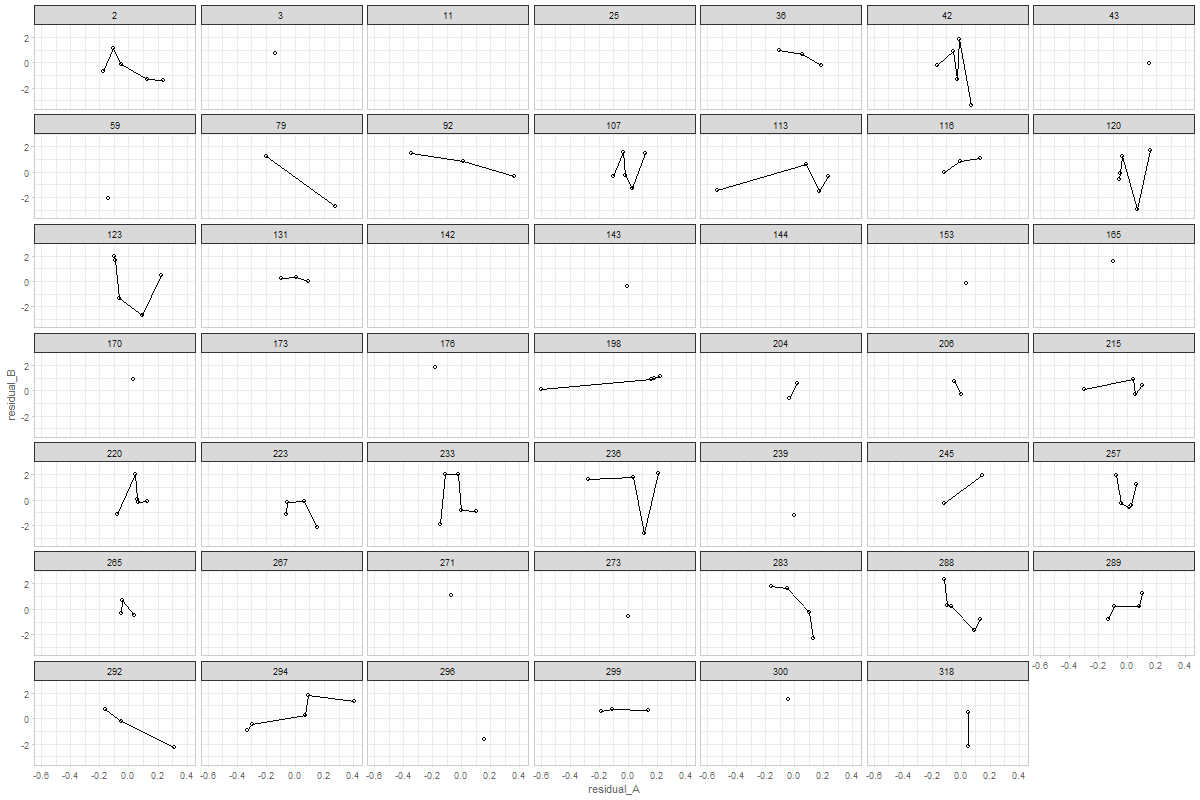
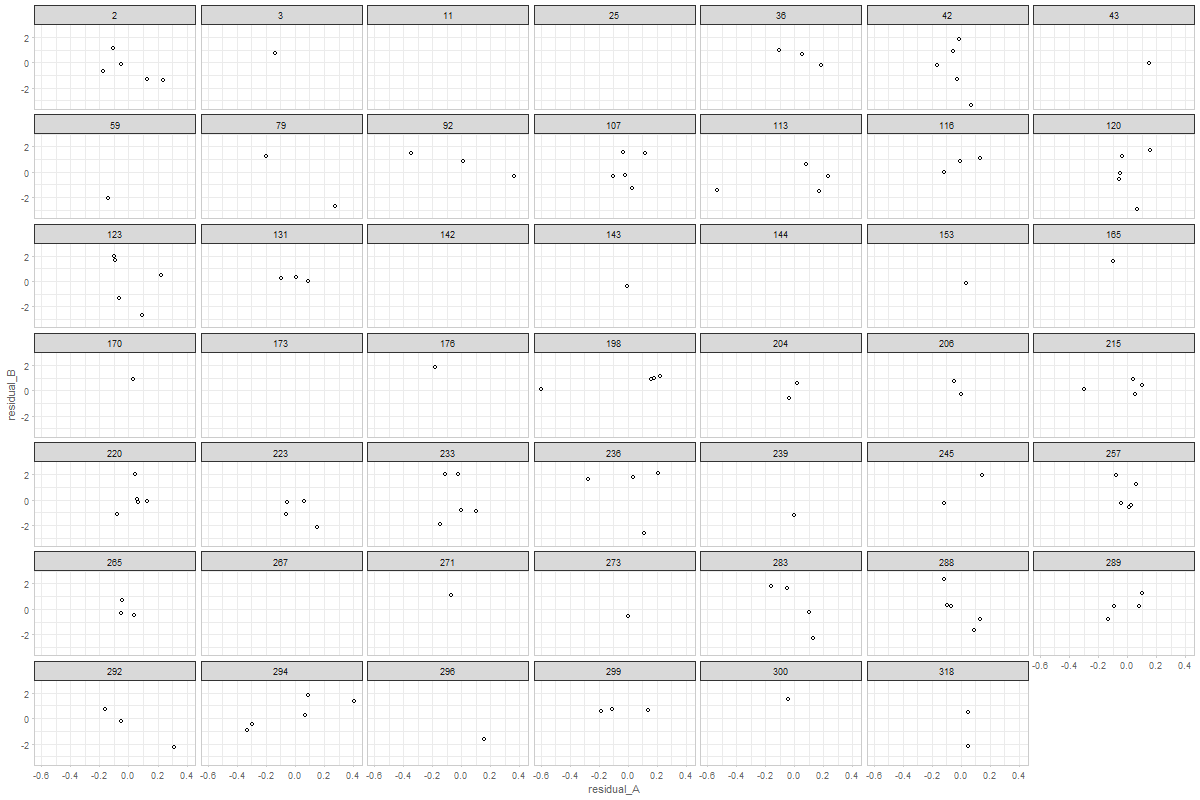 The graph shows a random 48 ids from MAP study.
The graph shows a random 48 ids from MAP study.
would it make sense to connect the dots?
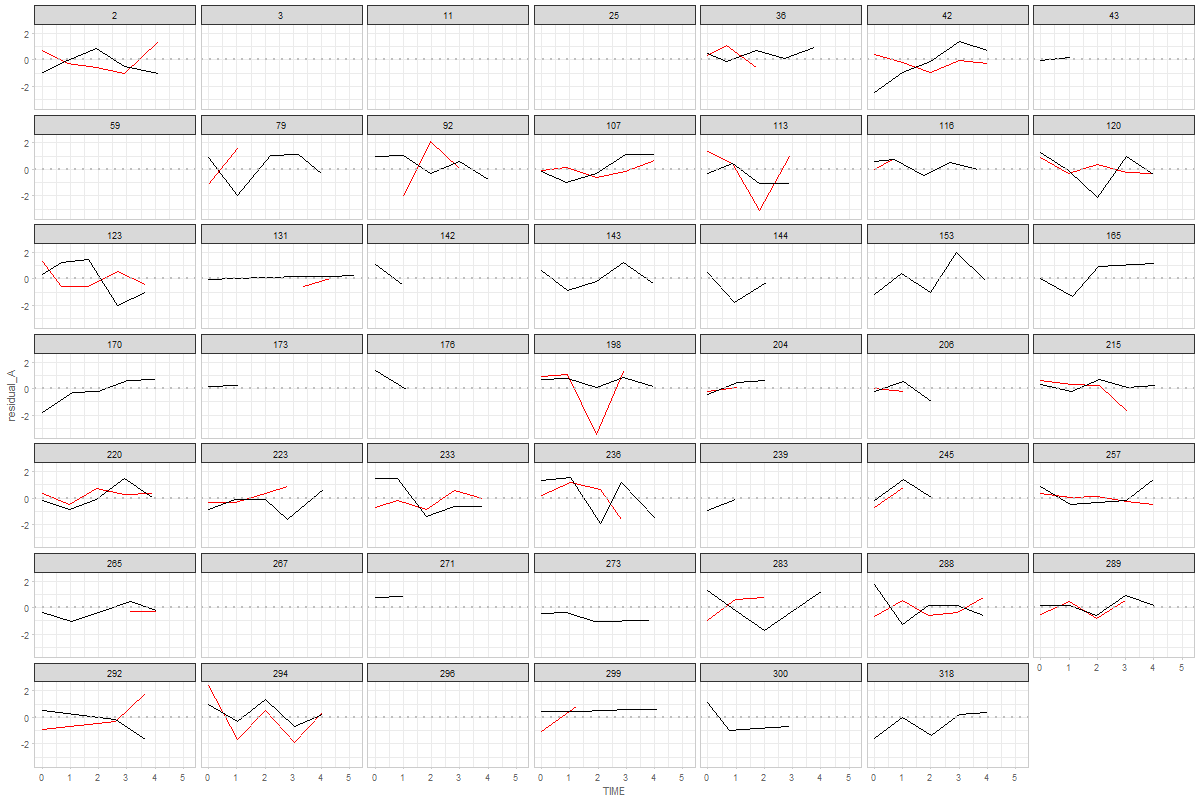
here's a version of this graph with connected dots.
You can adjust or further develop graph in the ./sandbox/coupling-graphs/coupling-graphs-1.R script. I have prepared the functions to extract the data. Let me know if you have any questions!
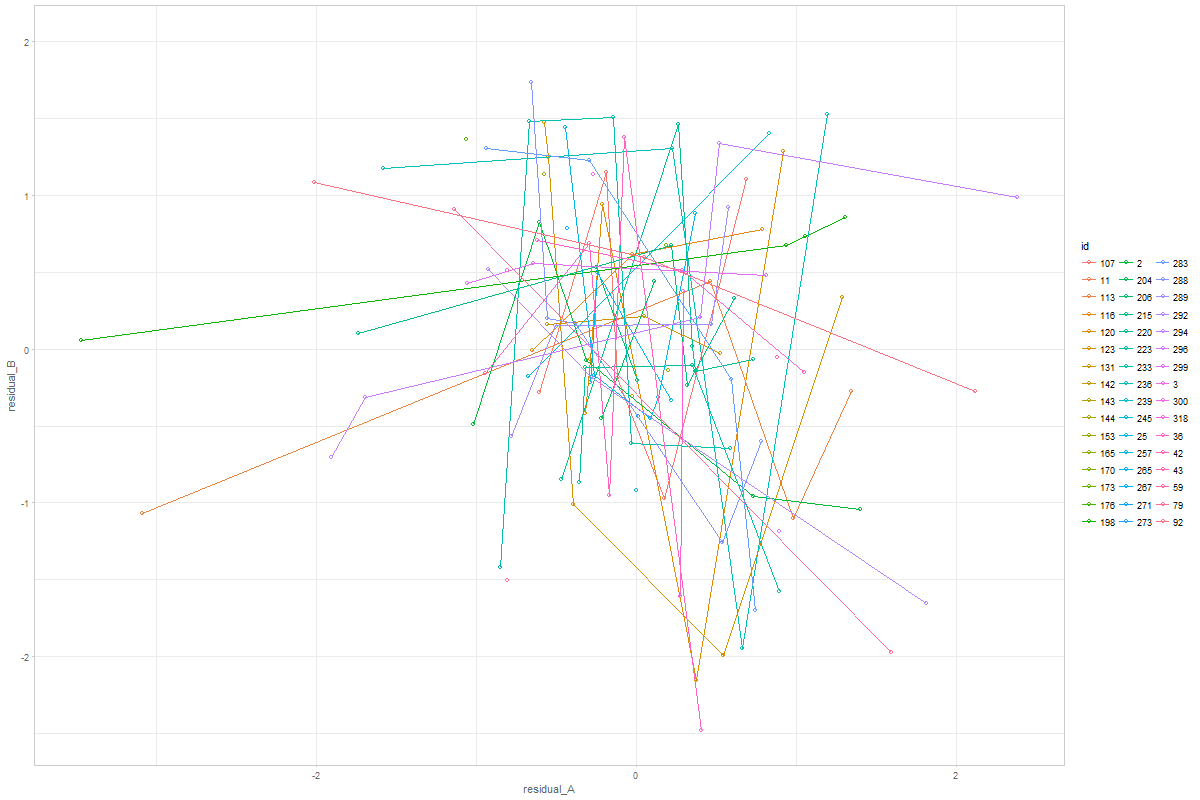
@ampiccinin @knighttime Objective: