OCR Visual Tables into Pandas DataFrames from PDF/DOC(X)/PPT files, 1000+ new state-of-the-art transformer models for Question Answering (QA) for over 30 languages, up to 700% speedup on GPU, 20 Biomedical models for over 8 languages, 50+ Terminology Code Mappers between RXNORM, NDC, UMLS,ICD10, ICDO, UMLS, SNOMED and MESH, Deidentification in Romanian, various Spark NLP helper methods and much more in 1 line of code with John Snow Labs NLU 4.0.0
NLU 4.0 for OCR Overview
On the OCR side, we now support extracting tables from PDF/DOC(X)/PPT files into structured pandas dataframe, making it easier than ever before to analyze bulks of files visually!
On the NLU core side we have over 1000+ new state-of-the-art models in over 30 languages for modern extractive transformer-based Question Answering problems powerd by the ALBERT/BERT/DistilBERT/DeBERTa/RoBERTa/Longformer Spark NLP Annotators trained on various SQUAD-like QA datasets for domains like Twitter, Tech, News, Biomedical COVID-19 and in various model subflavors like sci_bert, electra, mini_lm, covid_bert, bio_bert, indo_bert, muril, sapbert, bioformer, link_bert, mac_bert
Additionally up to 700% speedup transformer-based Word Embeddings on GPU and up to 97% speedup on CPU for tensorflow operations, support for Apple M1 chips, Pyspark 3.2 and 3.3 support.
Ontop of this, we are now supporting Apple M1 based architectures and every Pyspark 3.X version, while deprecating support for Pyspark 2.X.
Finally, NLU-Core features various new helper methods for working with Spark NLP and embellishes now the entire universe of Annotators defined by Spark NLP and Spark NLP for healthcare.
NLU 4.0 for Healthcare Overview
On the healthcare side NLU features 20 Biomedical models for over 8 languages (English, French, Italian, Portuguese, Romanian, Catalan and Galician) detect entities like HUMAN and SPECIES based on LivingNER corpus
Romanian models for Deidentification and extracting Medical entities like Measurements, Form, Symptom, Route, Procedure, Disease_Syndrome_Disorder, Score, Drug_Ingredient, Pulse, Frequency, Date, Body_Part, Drug_Brand_Name, Time, Direction, Dosage, Medical_Device, Imaging_Technique, Test, Imaging_Findings, Imaging_Test, Test_Result, Weight, Clinical_Dept and Units with SPELL and SPELL respectively
English NER models for parsing entities in Clinical Trial Abstracts like Age, AllocationRatio, Author, BioAndMedicalUnit, CTAnalysisApproach, CTDesign, Confidence, Country, DisorderOrSyndrome, DoseValue, Drug, DrugTime, Duration, Journal, NumberPatients, PMID, PValue, PercentagePatients, PublicationYear, TimePoint, Value using en.med_ner.clinical_trials_abstracts.pipe and also Pathogen NER models for Pathogen, MedicalCondition, Medicine with en.med_ner.pathogen and GENE_PROTEIN with en.med_ner.biomedical_bc2gm.pipeline
First Public Health Model for Emotional Stress classification It is a PHS-BERT-based model and trained with the Dreaddit dataset using en.classify.stress
50 + new Entity Mappers for problems like :
Extract section headers in scientific articles and normalize them with en.map_entity.section_headers_normalized
Map medical abbreviates to their definitions with en.map_entity.abbreviation_to_definition
Map drugs to action and treatments with en.map_entity.drug_to_action_treatment
Map drug brand to their National Drug Code (NDC) with en.map_entity.drug_brand_to_ndc
Convert between terminologies using en.<START_TERMINOLOGY>_to_<TARGET_TERMINOLOGY>
This works for the terminologies rxnorm, ndc, umls, icd10cm, icdo, umls, snomed, mesh
snomed_to_icdo
snomed_to_icd10cm
rxnorm_to_umls
powerd by Spark NLP for Healthcares ChunkMapper Annotator
Extract Tables from PDF files as Pandas DataFrames
Albert, Bert, DeBerta, DistilBert, LongFormer, RoBerta, XlmRoBerta based Transformer Architectures are now avaiable for question answering with almost 1000 models avaiable for 35 unique languages powerd by their corrosponding Spark NLP XXXForQuestionAnswering Annotator Classes and in various tuning and dataset flavours.
<lang>.answer_question.<domain>.<datasets>.<annotator_class><tune info>.by_<username>
If multiple datasets or tune parameters are defined , they are connected with a _ .
These substrings define up the <domain> part of the NLU reference
You need to use one of the Data formats below to pass context and question correctly to the model.
# use ||| to seperate question||context
data = 'What is my name?|||My name is Clara and I live in Berkeley'
# pass a tuple (question,context)
data = ('What is my name?','My name is Clara and I live in Berkeley')
# use pandas Dataframe, one column = question, one column=context
data = pd.DataFrame({
'question': ['What is my name?'],
'context': ["My name is Clara and I live in Berkely"]
})
# Get your answers with any of above formats
nlu.load("en.answer_question.squadv2.deberta").predict(data)
Visualize input data with an already configured Spark NLP pipeline,
for Algorithms of type (Ner,Assertion, Relation, Resolution, Dependency)
using Spark NLP Display
Automatically infers applicable viz type and output columns to use for visualization.
Example:
# works with Pipeline, LightPipeline, PipelineModel,PretrainedPipeline List[Annotator]
ade_pipeline = PretrainedPipeline('explain_clinical_doc_ade', 'en', 'clinical/models')
text = """I have an allergic reaction to vancomycin.
My skin has be itchy, sore throat/burning/itchy, and numbness in tongue and gums.
I would not recommend this drug to anyone, especially since I have never had such an adverse reaction to any other medication."""
nlu.viz(ade_pipeline, text)
returns:
If a pipeline has multiple models candidates that can be used for a viz,
the first Annotator that is vizzable will be used to create viz.
You can specify which type of viz to create with the viz_type parameter
Output columns to use for the viz are automatically deducted from the pipeline, by using the
first annotator that provides the correct output type for a specific viz.
You can specify which columns to use for a viz by using the
corresponding ner_col, pos_col, dep_untyped_col, dep_typed_col, resolution_col, relation_col, assertion_col, parameters.
nlu.autocomplete_pipeline(pipe)
Auto-Complete a pipeline or single annotator into a runnable pipeline by harnessing NLU's DAG Autocompletion algorithm and returns it as NLU pipeline.
The standard Spark pipeline is avaiable on the .vanilla_transformer_pipe attribute of the returned nlu pipe
Every Annotator and Pipeline of Annotators defines a DAG of tasks, with various dependencies that must be satisfied in topoligical order.
NLU enables the completion of an incomplete DAG by finding or creating a path between
the very first input node which is almost always is DocumentAssembler/MultiDocumentAssembler
and the very last node(s), which is given by the topoligical sorting the iterable annotators parameter.
Paths are created by resolving input features of annotators to the corrrosponding providers with matching storage references.
Example:
# Lets autocomplete the pipeline for a RelationExtractionModel, which as many input columns and sub-dependencies.
from sparknlp_jsl.annotator import RelationExtractionModel
re_model = RelationExtractionModel().pretrained("re_ade_clinical", "en", 'clinical/models').setOutputCol('relation')
text = """I have an allergic reaction to vancomycin.
My skin has be itchy, sore throat/burning/itchy, and numbness in tongue and gums.
I would not recommend this drug to anyone, especially since I have never had such an adverse reaction to any other medication."""
nlu_pipe = nlu.autocomplete_pipeline(re_model)
nlu_pipe.predict(text)
returns :
relation
relation_confidence
relation_entity1
relation_entity2
relation_entity2_class
1
1
allergic reaction
vancomycin
Drug_Ingredient
1
1
skin
itchy
Symptom
1
0.99998
skin
sore throat/burning/itchy
Symptom
1
0.956225
skin
numbness
Symptom
1
0.999092
skin
tongue
External_body_part_or_region
0
0.942927
skin
gums
External_body_part_or_region
1
0.806327
itchy
sore throat/burning/itchy
Symptom
1
0.526163
itchy
numbness
Symptom
1
0.999947
itchy
tongue
External_body_part_or_region
0
0.994618
itchy
gums
External_body_part_or_region
0
0.994162
sore throat/burning/itchy
numbness
Symptom
1
0.989304
sore throat/burning/itchy
tongue
External_body_part_or_region
0
0.999969
sore throat/burning/itchy
gums
External_body_part_or_region
1
1
numbness
tongue
External_body_part_or_region
1
1
numbness
gums
External_body_part_or_region
1
1
tongue
gums
External_body_part_or_region
nlu.to_pretty_df(pipe,data)
Annotates a Pandas Dataframe/Pandas Series/Numpy Array/Spark DataFrame/Python List strings /Python String
with given Spark NLP pipeline, which is assumed to be complete and runnable and returns it in a pythonic pandas dataframe format.
Example:
# works with Pipeline, LightPipeline, PipelineModel,PretrainedPipeline List[Annotator]
ade_pipeline = PretrainedPipeline('explain_clinical_doc_ade', 'en', 'clinical/models')
text = """I have an allergic reaction to vancomycin.
My skin has be itchy, sore throat/burning/itchy, and numbness in tongue and gums.
I would not recommend this drug to anyone, especially since I have never had such an adverse reaction to any other medication."""
# output is same as nlu.autocomplete_pipeline(re_model).nlu_pipe.predict(text)
nlu.to_pretty_df(ade_pipeline,text)
returns :
assertion
asserted_entitiy
entitiy_class
assertion_confidence
present
allergic reaction
ADE
0.998
present
itchy
ADE
0.8414
present
sore throat/burning/itchy
ADE
0.9019
present
numbness in tongue and gums
ADE
0.9991
Annotators are grouped internally by NLU into output levels token,sentence, document,chunk and relation
Same level annotators output columns are zipped and exploded together to create the final output df.
Additionally, most keys from the metadata dictionary in the result annotations will be collected and expanded into their own columns in the resulting Dataframe, with special handling for Annotators that encode multiple metadata fields inside of one, seperated by strings like ||| or :::.
Some columns are omitted from metadata to reduce total amount of output columns, these can be re-enabled by setting metadata=True
For a given pipeline output level is automatically set to the last anntators output level by default.
This can be changed by defining to_preddty_df(pipe,text,output_level='my_level' for levels token,sentence, document,chunk and relation .
nlu.to_nlu_pipe(pipe)
Convert a pipeline or list of annotators into a NLU pipeline making .predict() and .viz() avaiable for every Spark NLP pipeline.
Assumes the pipeline is already runnable.
# works with Pipeline, LightPipeline, PipelineModel,PretrainedPipeline List[Annotator]
ade_pipeline = PretrainedPipeline('explain_clinical_doc_ade', 'en', 'clinical/models')
text = """I have an allergic reaction to vancomycin.
My skin has be itchy, sore throat/burning/itchy, and numbness in tongue and gums.
I would not recommend this drug to anyone, especially since I have never had such an adverse reaction to any other medication."""
nlu_pipe = nlu.to_nlu_pipe(ade_pipeline)
# Same output as nlu.to_pretty_df(pipe,text)
nlu_pipe.predict(text)
# same output as nlu.viz(pipe,text)
nlu_pipe.viz(text)
# Acces auto-completed Spark NLP big data pipeline,
nlu_pipe.vanilla_transformer_pipe.transform(spark_df)
returns :
assertion
asserted_entitiy
entitiy_class
assertion_confidence
present
allergic reaction
ADE
0.998
present
itchy
ADE
0.8414
present
sore throat/burning/itchy
ADE
0.9019
present
numbness in tongue and gums
ADE
0.9991
and
4 new Demo Notebooks
These notebooks showcase some of latest classifier models for Banking Queries, Intents in Text, Question and new s classification
NLU captures every Annotator of Spark NLP and Spark NLP for healthcare
The entire universe of Annotators in Spark NLP and Spark-NLP for healthcare is now embellished by NLU Components by using generalizable annotation extractors methods and configs internally to support enable the new NLU util methods.
The following annotator classes are newly captured:
nlu.load('en.ner.clinical_trials_abstracts').predict('A one-year, randomised, multicentre trial comparing insulin glargine with NPH insulin in combination with oral agents in patients with type 2 diabetes.')
Results:
entities_clinical_trials_abstracts
entities_clinical_trials_abstracts_class
entities_clinical_trials_abstracts_confidence
0
randomised
CTDesign
0.9996
0
multicentre
CTDesign
0.9998
0
insulin glargine
Drug
0.99135
0
NPH insulin
Drug
0.96875
0
type 2 diabetes
DisorderOrSyndrome
0.999933
Code:
nlu.load('en.ner.clinical_trials_abstracts').viz('A one-year, randomised, multicentre trial comparing insulin glargine with NPH insulin in combination with oral agents in patients with type 2 diabetes.')
nlu.load('en.med_ner.pathogen').predict('Racecadotril is an antisecretory medication and it has better tolerability than loperamide. Diarrhea is the condition of having loose, liquid or watery bowel movements each day. Signs of dehydration often begin with loss of the normal stretchiness of the skin. While it has been speculated that rabies virus, Lyssavirus and Ephemerovirus could be transmitted through aerosols, studies have concluded that this is only feasible in limited conditions.')
Results:
entities_pathogen
entities_pathogen_class
entities_pathogen_confidence
0
Racecadotril
Medicine
0.9468
0
loperamide
Medicine
0.9987
0
Diarrhea
MedicalCondition
0.9848
0
dehydration
MedicalCondition
0.6307
0
rabies virus
Pathogen
0.95685
0
Lyssavirus
Pathogen
0.9694
0
Ephemerovirus
Pathogen
0.6917
Code:
nlu.load('en.med_ner.pathogen').viz('Racecadotril is an antisecretory medication and it has better tolerability than loperamide. Diarrhea is the condition of having loose, liquid or watery bowel movements each day. Signs of dehydration often begin with loss of the normal stretchiness of the skin. While it has been speculated that rabies virus, Lyssavirus and Ephemerovirus could be transmitted through aerosols, studies have concluded that this is only feasible in limited conditions.')
nlu.load('es.med_ner.living_species.roberta').predict('Lactante varón de dos años. Antecedentes familiares sin interés. Antecedentes personales: Embarazo, parto y periodo neonatal normal. En seguimiento por alergia a legumbres, diagnosticado con diez meses por reacción urticarial generalizada con lentejas y garbanzos, con dieta de exclusión a legumbres desde entonces. En ésta visita la madre describe episodios de eritema en zona maxilar derecha con afectación ocular ipsilateral que se resuelve en horas tras la administración de corticoides. Le ha ocurrido en 5-6 ocasiones, en relación con la ingesta de alimentos previamente tolerados. Exploración complementaria: Cacahuete, ac(ige)19.2 Ku.arb/l. Resultados: Ante la sospecha clínica de Síndrome de Frey, se tranquiliza a los padres, explicándoles la naturaleza del cuadro y se cita para revisión anual.')
Results:
entities_living_species
entities_living_species_class
entities_living_species_confidence
0
Lactante varón
HUMAN
0.93175
0
familiares
HUMAN
1
0
personales
HUMAN
1
0
neonatal
HUMAN
0.9997
0
legumbres
SPECIES
0.9962
0
lentejas
SPECIES
0.9988
0
garbanzos
SPECIES
0.9901
0
legumbres
SPECIES
0.9976
0
madre
HUMAN
1
0
Cacahuete
SPECIES
0.998
0
padres
HUMAN
1
Code:
nlu.load('es.med_ner.living_species.roberta').viz('Lactante varón de dos años. Antecedentes familiares sin interés. Antecedentes personales: Embarazo, parto y periodo neonatal normal. En seguimiento por alergia a legumbres, diagnosticado con diez meses por reacción urticarial generalizada con lentejas y garbanzos, con dieta de exclusión a legumbres desde entonces. En ésta visita la madre describe episodios de eritema en zona maxilar derecha con afectación ocular ipsilateral que se resuelve en horas tras la administración de corticoides. Le ha ocurrido en 5-6 ocasiones, en relación con la ingesta de alimentos previamente tolerados. Exploración complementaria: Cacahuete, ac(ige)19.2 Ku.arb/l. Resultados: Ante la sospecha clínica de Síndrome de Frey, se tranquiliza a los padres, explicándoles la naturaleza del cuadro y se cita para revisión anual.')
OCR Visual Tables into Pandas DataFrames from PDF/DOC(X)/PPT files, 1000+ new state-of-the-art transformer models for Question Answering (QA) for over 30 languages, up to 700% speedup on GPU, 20 Biomedical models for over 8 languages, 50+ Terminology Code Mappers between RXNORM, NDC, UMLS,ICD10, ICDO, UMLS, SNOMED and MESH, Deidentification in Romanian, various Spark NLP helper methods and much more in 1 line of code with John Snow Labs NLU 4.0.0
NLU 4.0 for OCR Overview
On the OCR side, we now support extracting tables from PDF/DOC(X)/PPT files into structured pandas dataframe, making it easier than ever before to analyze bulks of files visually!
Checkout the OCR Tutorial for extracting to see this in action
to see this in action
Tablesfrom Image/PDF/DOC(X) filesThese models grab all Table data from the files detected and return a
list of Pandas DataFrames,containing Pandas DataFrame for every table detected
pdf2table)ppt2table)doc2table)This is powerd by John Snow Labs Spark OCR Annotataors for PdfToTextTable, DocToTextTable, PptToTextTable
NLU 4.0 Core Overview
On the NLU core side we have over 1000+ new state-of-the-art models in over 30 languages for modern extractive transformer-based Question Answering problems powerd by the ALBERT/BERT/DistilBERT/DeBERTa/RoBERTa/Longformer Spark NLP Annotators trained on various SQUAD-like QA datasets for domains like Twitter, Tech, News, Biomedical COVID-19 and in various model subflavors like sci_bert, electra, mini_lm, covid_bert, bio_bert, indo_bert, muril, sapbert, bioformer, link_bert, mac_bert
Additionally up to 700% speedup transformer-based Word Embeddings on GPU and up to 97% speedup on CPU for tensorflow operations, support for Apple M1 chips, Pyspark 3.2 and 3.3 support. Ontop of this, we are now supporting Apple M1 based architectures and every Pyspark 3.X version, while deprecating support for Pyspark 2.X.
Finally, NLU-Core features various new helper methods for working with Spark NLP and embellishes now the entire universe of Annotators defined by Spark NLP and Spark NLP for healthcare.
NLU 4.0 for Healthcare Overview
On the healthcare side NLU features 20 Biomedical models for over 8 languages (English, French, Italian, Portuguese, Romanian, Catalan and Galician) detect entities like
HUMANandSPECIESbased on LivingNER corpusRomanian models for Deidentification and extracting Medical entities like
Measurements,Form,Symptom,Route,Procedure,Disease_Syndrome_Disorder,Score,Drug_Ingredient,Pulse,Frequency,Date,Body_Part,Drug_Brand_Name,Time,Direction,Dosage,Medical_Device,Imaging_Technique,Test,Imaging_Findings,Imaging_Test,Test_Result,Weight,Clinical_DeptandUnitswith SPELL and SPELL respectivelyEnglish NER models for parsing entities in Clinical Trial Abstracts like
Age,AllocationRatio,Author,BioAndMedicalUnit,CTAnalysisApproach,CTDesign,Confidence,Country,DisorderOrSyndrome,DoseValue,Drug,DrugTime,Duration,Journal,NumberPatients,PMID,PValue,PercentagePatients,PublicationYear,TimePoint,Valueusingen.med_ner.clinical_trials_abstracts.pipeand also Pathogen NER models forPathogen,MedicalCondition,Medicinewithen.med_ner.pathogenandGENE_PROTEINwithen.med_ner.biomedical_bc2gm.pipelineFirst Public Health Model for Emotional Stress classification It is a PHS-BERT-based model and trained with the Dreaddit dataset using
en.classify.stress50 + new Entity Mappers for problems like :
en.map_entity.section_headers_normalizeden.map_entity.abbreviation_to_definitionen.map_entity.drug_to_action_treatmenten.map_entity.drug_brand_to_ndcen.<START_TERMINOLOGY>_to_<TARGET_TERMINOLOGY>rxnorm,ndc,umls,icd10cm,icdo,umls,snomed,meshsnomed_to_icdosnomed_to_icd10cmrxnorm_to_umlsExtract Tables from PDF files as Pandas DataFrames
Sample PDF: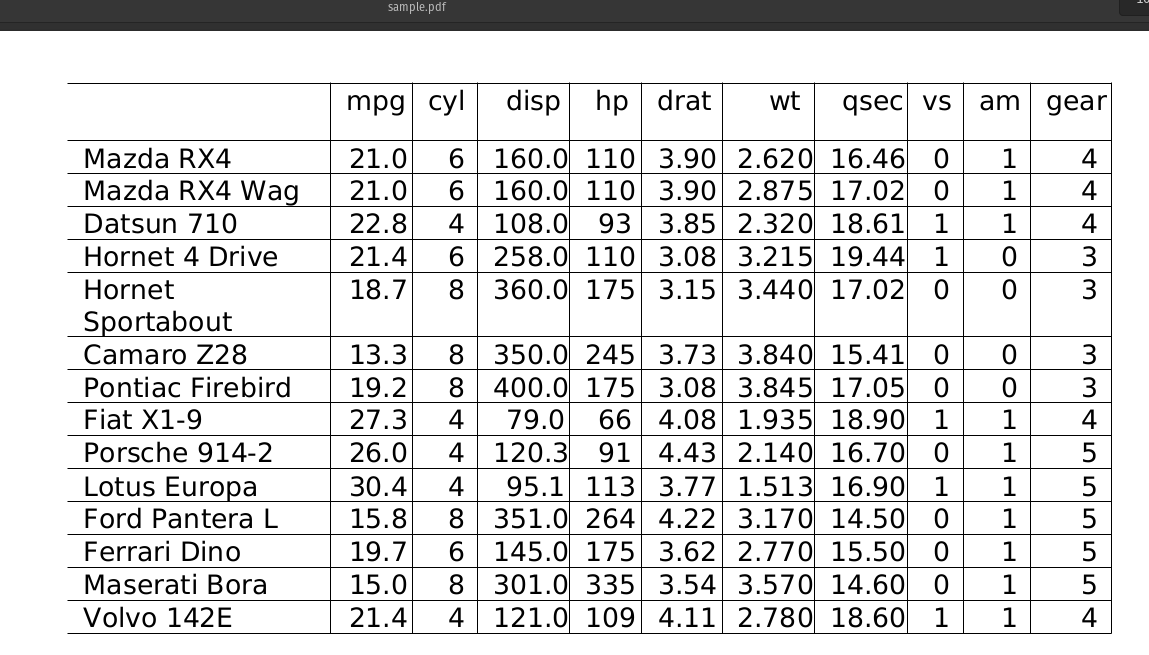
Output of PDF Table OCR :
Extract Tables from DOC/DOCX files as Pandas DataFrames
Sample DOCX: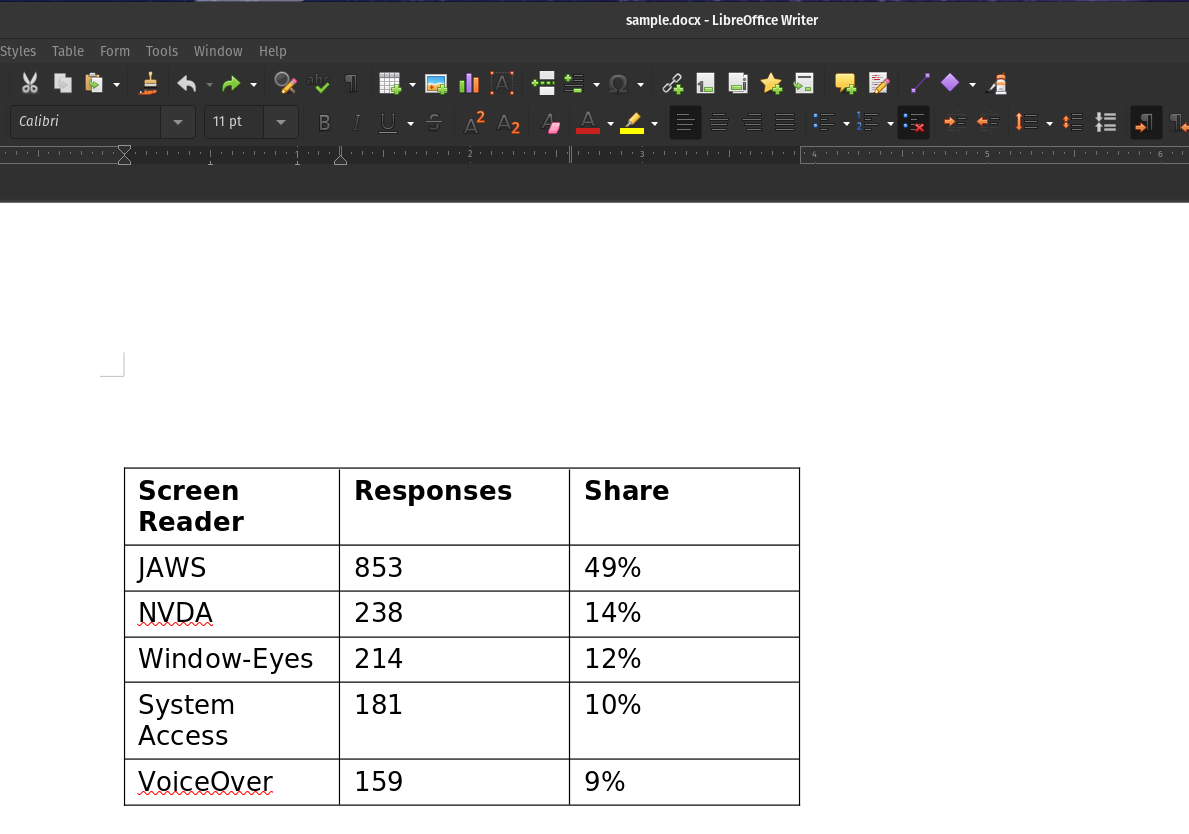
Output of DOCX Table OCR :
Extract Tables from PPT files as Pandas DataFrame
Sample PPT with two tables:
Output of PPT Table OCR :
and
Span Classifiers for question answering
Albert, Bert, DeBerta, DistilBert, LongFormer, RoBerta, XlmRoBerta based Transformer Architectures are now avaiable for question answering with almost 1000 models avaiable for 35 unique languages powerd by their corrosponding Spark NLP XXXForQuestionAnswering Annotator Classes and in various tuning and dataset flavours.
<lang>.answer_question.<domain>.<datasets>.<annotator_class><tune info>.by_<username>If multiple datasets or tune parameters are defined , they are connected with a_.These substrings define up the
<domain>part of the NLU referenceThese substrings define up the
<dataset>part of the NLU referenceThese substrings define up the
<dataset>part of the NLU referenceThese substrings define the
<annotator_class>substring, if it does not map to a sparknlp annotatorThese substrings define the
<tune_info>substring, if it does not map to a sparknlp annotatormultilingual,mini_lm,xtremedistiled,distilled,xtreme,augmented,zero_shotxl,xxl,large,base,medium,base,small,tiny,cased,uncased1024d,768d,512d,256d,128d,64d,32dQA DataFormat
You need to use one of the Data formats below to pass context and question correctly to the model.
returns :
New NLU helper Methods
You can see all features showcased in the notebook or on the new docs page for Spark NLP utils
notebook or on the new docs page for Spark NLP utils
nlu.viz(pipe,data)
Visualize input data with an already configured Spark NLP pipeline,
for Algorithms of type (Ner,Assertion, Relation, Resolution, Dependency)
using Spark NLP Display
Automatically infers applicable viz type and output columns to use for visualization.
Example:
returns:
If a pipeline has multiple models candidates that can be used for a viz,
the first Annotator that is vizzable will be used to create viz.
You can specify which type of viz to create with the viz_type parameter
Output columns to use for the viz are automatically deducted from the pipeline, by using the first annotator that provides the correct output type for a specific viz.
You can specify which columns to use for a viz by using the
corresponding ner_col, pos_col, dep_untyped_col, dep_typed_col, resolution_col, relation_col, assertion_col, parameters.
nlu.autocomplete_pipeline(pipe)
Auto-Complete a pipeline or single annotator into a runnable pipeline by harnessing NLU's DAG Autocompletion algorithm and returns it as NLU pipeline. The standard Spark pipeline is avaiable on the
.vanilla_transformer_pipeattribute of the returned nlu pipeEvery Annotator and Pipeline of Annotators defines a
DAGof tasks, with various dependencies that must be satisfied intopoligical order. NLU enables the completion of an incomplete DAG by finding or creating a path between the very first input node which is almost always isDocumentAssembler/MultiDocumentAssemblerand the very last node(s), which is given by thetopoligical sortingthe iterable annotators parameter. Paths are created by resolving input features of annotators to the corrrosponding providers with matching storage references.Example:
returns :
nlu.to_pretty_df(pipe,data)
Annotates a Pandas Dataframe/Pandas Series/Numpy Array/Spark DataFrame/Python List strings /Python String
with given Spark NLP pipeline, which is assumed to be complete and runnable and returns it in a pythonic pandas dataframe format.
Example:
returns :
Annotators are grouped internally by NLU into output levels
token,sentence,document,chunkandrelationSame level annotators output columns are zipped and exploded together to create the final output df. Additionally, most keys from the metadata dictionary in the result annotations will be collected and expanded into their own columns in the resulting Dataframe, with special handling for Annotators that encode multiple metadata fields inside of one, seperated by strings like|||or:::. Some columns are omitted from metadata to reduce total amount of output columns, these can be re-enabled by settingmetadata=TrueFor a given pipeline output level is automatically set to the last anntators output level by default. This can be changed by defining
to_preddty_df(pipe,text,output_level='my_level'for levelstoken,sentence,document,chunkandrelation.nlu.to_nlu_pipe(pipe)
Convert a pipeline or list of annotators into a NLU pipeline making
.predict()and.viz()avaiable for every Spark NLP pipeline. Assumes the pipeline is already runnable.returns :
and
4 new Demo Notebooks
These notebooks showcase some of latest classifier models for Banking Queries, Intents in Text, Question and new s classification
NLU captures every Annotator of Spark NLP and Spark NLP for healthcare
The entire universe of Annotators in Spark NLP and Spark-NLP for healthcare is now embellished by NLU Components by using generalizable annotation extractors methods and configs internally to support enable the new NLU util methods. The following annotator classes are newly captured:
All NLU 4.0 for Healthcare Models
Some examples:
en.rxnorm.umls.mapping
Code:
en.ner.clinical_trials_abstracts
Code:
Results:
Code:
Results:
en.med_ner.pathogen
Code:
Results:
Code:
Results:
es.med_ner.living_species.roberta
Code:
Results:
Code:
Results:
All healthcare models added in NLU 4.0 :
All NLU 4.0 Core Models
All core models added in NLU 4.0 : Can be found on the NLU website because of Github Limitations