Hi -- happy to help.
The way you would do that with AbstractGPs.jl / Stheno.jl is something like:
x = range(0.0, 1.0; length=1_000)
f = GP(SEKernel() ∘ ScaleTransform(1 / 0.2))
sampleplot(f(x); samples=10)(I might be off by a factor of two with the length scale or something)
Internally, kernelmatrix(SEKernel() ∘ ScaleTransform(1 / 0.2), x) will be called, which is what I think you're after with K(x).

I've been trying to figure out how to utilize Stheno.jl to duplicate some code I found in python. I'm new to kernels and how they are used so don't hesitate to explain what I'm not understanding. The python code uses the squared exponential kernel (Radial Basis Functions) to produce n random but smooth functions:
When I plot the results I get plots that look like the following for n=10: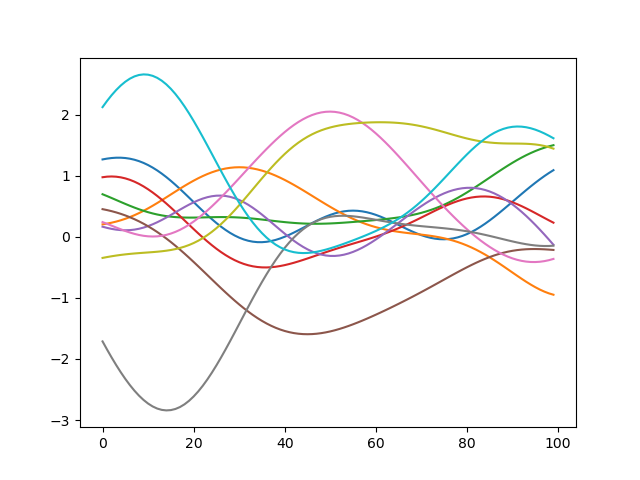
Unfortunately I get stuck early on when I want to produce the radial basis functions of x (
k=K(x)). Any help is appreciated.