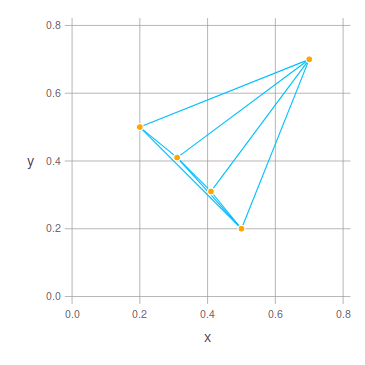
I realised that in a last-minute attempt to simplify the code I introduced a bug in the circumcircleUnion function.
In the previous commit, for each edge of the convex hull the circumcircles of the edge and any of the other points is computed. Then the circumcircle with the largest radius was selected and added to the union of circumcircles. This does not result in the correct union of circumcircles, because the selected circumcircle is often rather in the inner of the convex hull of points and circumcircles that extend the most towards the outside of the convex hull are missed.
The first plot shows the output of the corrected function circumcircleUnion in green. For each edge of the convex hull, the function starts off with the circumcircle of the edge. It then checks all the other points from the point set that happen to lie in the circumcircle and computes the circumcircle of the point and the edge points - if its radius is larger than the radius of the previous circumcircle, then the circumcircle is updated. By restriction to the points inside the circumcircle we can only obtain circumcircles that cover a larger region outside the convex hull than the "edge circumcircle". If we take the union of the computed set and the inner of the convex hull of points, we end up with a superset of the actual union of the circumcircles of the triangles of the Delaunay tessellation, which is sufficient to ensure convexity of the scaled tessellation. The second plot shows the final tessellation.





Closes #34 and #27.
Non-convexity of the tessellation means we are missing triangles. A non-convex tessellation is obtained, if at least one of the corner points of the initial square lies inside the circumcircle of a missing triangle. This problem is solved by scaling and shifting the points such that the union of the circumcircles of the triangles along the boundary of the tessellation lies inside (1,2)x(1,2). With this modification it is now possible to start off with a point set outside [1,2]x[1,2]. Apply the function scaleShiftPoints to the point set before the tessellation and the function expand to the edge vertices after the tessellation.
Here's an example:
This produces the following plot: