Dear Harish, her some thoughts in a rush to your questions. Hope I understood you correct.
1) you have to add the CYP3A4 enzyme to your individual BB. Easiest way would be to use the PK-Sim expression database. However there is only one for human available. Nevertheless you can yous this as a starting point for rats. It will give you the CYP3A4 abundance for humans. You can use rat values as well if you have. For details how to use it check out here: https://github.com/Open-Systems-Pharmacology/Forum/issues/279
2) You can input this Clint values in the compound BB, ADME properties Tab. You have to create a metabolic process for your desired enzymes. Again for details check the videos.
3) your t1/2 in hepatocytes can be inputted in the compound BB as total hepatic clearance. Select input from half live in hepatocytes. Be sure to provide the correct number of hepatocytes you used in your assay. 4) in the compound BB, ADME properties Tab you can also input renal clearance e.g. as renal plasma clearance or active secretion process. Of course this can be fitted additional to a metabolic clearance as well.
B.tw.: your fit seems to describe the distribution of your compound very well. There seem to be some clearance process missing to catch the points after 2 hours. I think you are almost there. However, this is of course difficult to say without knowing your fitted parameters and without checking parameter estimation VPCs ;-)
Hope this might help you. Keep up the good work.
Best, Tobias
 This looks pretty good already. please note the fraction metabolised in liver and kidney and fraction excreted to urine. If you have measured counterparts in vivo you can use these for model qualification.
This looks pretty good already. please note the fraction metabolised in liver and kidney and fraction excreted to urine. If you have measured counterparts in vivo you can use these for model qualification.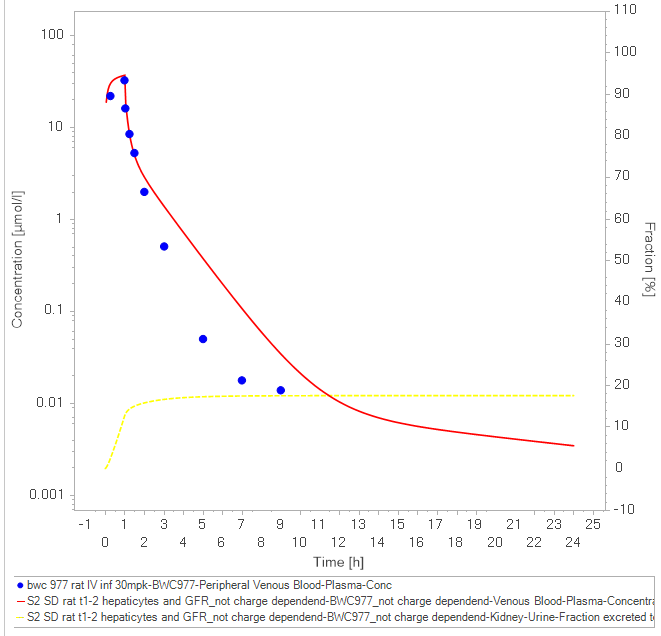 This looks not so bad, but could be a bit more CL. Please note the higher simulated
fraction renally excreted.
This looks not so bad, but could be a bit more CL. Please note the higher simulated
fraction renally excreted.
 Metabolism is probably underestimated here. Therefor I would go with S1. Again, if you have measurements for fraction renally excreted in rats you can use this to finally decide between the two model variants
Metabolism is probably underestimated here. Therefor I would go with S1. Again, if you have measurements for fraction renally excreted in rats you can use this to finally decide between the two model variants
Hello All, Iam trying to build PBPK model in rats for a NCE administered by IV Infusion for 1hour. I have physicochemical parameters, invitro data and invivo data. My questions are
In the individual building block apart from providing the biometrics , do we also have to provide enzyme expression (CYP3A4 etc)
I have converted CLint measured in hepatocytes through IVIVE and provided the Clint values for CYP3A4 enzyme
I have also provided t1/2 from hepatocytes data
Compound has liability for renal clearance so accounted for that by giving Renal process
Inspite of that fitting looks something like the attached figure. Any suggestions will be of great values as to what i have to keep in mind
Regards Harish