I think I found the data in the shared lab gDrive. In the 2016 AxonSeg paper, the CARS data is described as such:
CARS images were obtained from one rat (thoracic section). Sample was imaged with a 60× objective lens (UPLSAPO 1.2 NA w, Olympus) and recorded images were stitched to reconstruct the whole section (~0.2 μm/pixel).
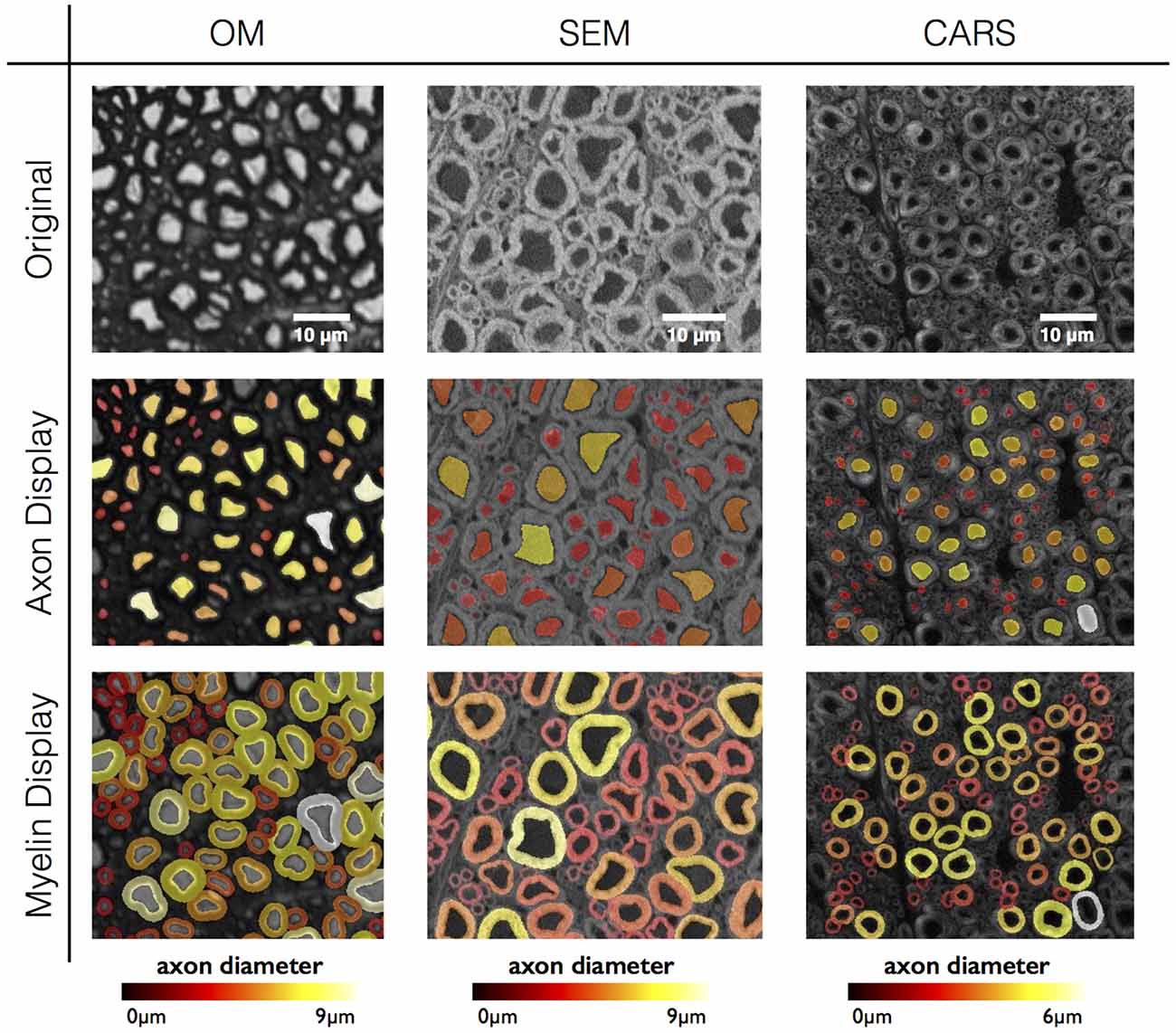
This is what the GT looks like:
Only problem is that only the axon masks are available as GT. The myelin segmentation was the result of another postprocessing step in this study, and only the axon segmentation was evaluated. With that being said, we should look for the myelin segmentations available and potentially use them as GTs. (some CARS myelin segs were shown in the paper so they obviously exist). I will dig deeper in duke but might run into perm issues.
We have annotated CARS data buried somewhere - deep within
dukeor something. We need to add this data to the study in order to cover all 4 major microscopy imaging modalities. I will update my progress for this task in this issue.