Looks like an issues with classes. Can you please provide a reproducible example (always do that if you can)?
Closed Steviey closed 2 years ago
Looks like an issues with classes. Can you please provide a reproducible example (always do that if you can)?
The reproduce able example works fine so far. Now I have a difference between the example code and a 30K-line-script :-). Which let me think, it could be related to the data-source. What does the cross in the graph mean? Noticed 1: Even LASSO and RIDGE are working now -not in the 'live-code'...have to investigate... Noticed 2: Changing the model-def can lead to failure because of src (normal ETS-behaviour) Noticed 3: Setting cumulative=FALSE leads to 'Failed to return valid external regressors.' src-variance issue (normal) Noticed 4: Shuffling the data can lead to the initial issue (example code updated).
options(scipen = 999)
options(dplyr.summarise.inform=F)
options(max.print=2000)
library("Hmisc")
suppressPackageStartupMessages(library(lightgbm))
suppressPackageStartupMessages(library(digest))
suppressPackageStartupMessages(library(RSQLite))
suppressPackageStartupMessages(library(stringr))
suppressPackageStartupMessages(library(tidyr))
suppressPackageStartupMessages(library(dbplyr))
suppressPackageStartupMessages(library(rlang))
suppressPackageStartupMessages(library(freqdist))
suppressPackageStartupMessages(library(tidymodels))
options(tidymodels.dark = TRUE)
suppressPackageStartupMessages(library(modeltime))
suppressPackageStartupMessages(library(modeltime.ensemble))
suppressPackageStartupMessages(library(modeltime.resample))
suppressPackageStartupMessages(library(timetk))
suppressPackageStartupMessages(library(tidyverse))
suppressPackageStartupMessages(library(rsample))
suppressPackageStartupMessages(library(tidyquant))
suppressPackageStartupMessages(library(tibbletime))
suppressPackageStartupMessages(library(anomalize))
suppressPackageStartupMessages(library(smooth))
suppressPackageStartupMessages(library(lmtest))
suppressPackageStartupMessages(library(mgcv))
suppressPackageStartupMessages(library(fable))
suppressPackageStartupMessages(library(fabletools))
suppressPackageStartupMessages(library(tsibble))
suppressPackageStartupMessages(library(tsibbledata))
suppressPackageStartupMessages(library(tsfeatures))
suppressPackageStartupMessages(library(ggplot2))
suppressPackageStartupMessages(library(ggrepel))
suppressPackageStartupMessages(library(runner))
suppressPackageStartupMessages(library(ggformula))
suppressPackageStartupMessages(library(fANCOVA))
suppressPackageStartupMessages(library(stats))
suppressPackageStartupMessages(library(TTR))
suppressPackageStartupMessages(library(xts))
suppressPackageStartupMessages(library(vip))
suppressPackageStartupMessages(library(yardstick))
suppressPackageStartupMessages(library(plotly))
suppressPackageStartupMessages(library(catboost))
suppressPackageStartupMessages(library(treesnip))
suppressPackageStartupMessages(library(broom))
suppressPackageStartupMessages(library(finetune))
suppressPackageStartupMessages(library(tabnet))
prepModteltimeData<-function(df){
ret<-list()
rownames(df) <- NULL
df <- df %>% dplyr::mutate(id=row_number()) %>% relocate(id)
ret$df <- df
df_tsbl <- df %>%
as_tsibble(index=id)
df_split <- rsample::initial_time_split(df_tsbl, prop = 0.8)
train_len <- length(df_split$in_id)
test_len <- length(df_split$out_id)
df_tsbl <- as.data.frame(df_tsbl) # modeltime input format
splits <- df_tsbl %>%
timetk::time_series_split(assess=test_len, cumulative = TRUE)#kann auch int rein!
ret$df_train <- rsample::training(splits)
ret$df_test <- rsample::testing(splits)
ret$df_split<-df_split
ret$ts_split<-splits
myH<-1
myLimit <- as.numeric(nrow(df_tsbl)-myH)
ret$trainOneStep <- df %>% filter(id <= myLimit)
ret$testOneStep <- df %>% filter(id > myLimit)
return(ret)
}
getFormulaLocalEn<-function(xRegCols){
xRegCols <- xRegCols[xRegCols!='value']
xRegCols <- xRegCols[xRegCols!='id']
xRegCols <- xRegCols[xRegCols!='date']
myFormula <- xRegCols
myFormula <- paste0(myFormula, collapse= "+")
myFormula <- paste0('value ~ date + ',myFormula)
myFormula <- as.formula(myFormula)
return(myFormula)
}
y<-rnorm(120, mean=15, sd=5)
x<-c(
rep(0.08,20)
,rep(0.20,20)
,rep(0.40,20)
,rep(0.60,20)
,rep(0.80,20)
,rep(1.0,20)
)
x1<-c(
rep(0.18,20)
,rep(0.21,20)
,rep(0.10,20)
,rep(0.61,20)
,rep(0.81,20)
,rep(1.0,20)
)
data <-data.frame(value=y,xReg=x,xReg1=x1)
# shuffle (new)
rows <-sample(nrow(data))
data <-data[rows, ]
dataLength <-nrow(data)
myTime <-tk_make_timeseries("2011", length_out=dataLength, include_endpoints = FALSE)
data$date <-myTime
dataObj <-prepModteltimeData(data)
df_train <-dataObj$df_train
df_test <-dataObj$df_test
allData <-dplyr::bind_rows(df_train,df_test)
df_split <-rsample::initial_time_split(df_train, prop = 0.75)
train_len <-length(df_split$in_id)
test_len <-length(df_split$out_id)
myFormula <-getFormulaLocalEn(c('xReg','xReg1'))
recipe_spec <- recipe(myFormula,df_train)
set.seed(123)
cvResamples <- allData %>%
time_series_cv(
assess = test_len
,initial = train_len
,slice_limit = 10
)
lossVect = c("RIDGE","LASSO","GTMSE","likelihood","MSE","MAE","HAM","MSEh","TMSE","MSCE")
algoGrid <- expand.grid(
lossParam=lossVect
,dummy=c('A')
,stringsAsFactors = FALSE
,KEEP.OUT.ATTRS = FALSE
)
results <- vector(mode = 'numeric', length = nrow(algoGrid))
idx<-0
for(ee in seq_len(nrow(algoGrid))){
params <- algoGrid[ee, ]
myLoss <- as.character(params[1,'lossParam'])
print(myLoss)
model_spec <- adam_reg(
ets_model = 'ANN'
,loss = myLoss
,distribution = 'ds'
) %>%
set_engine("adam",silent=FALSE,h=1)
model_spec<-parsnip::eval_args(model_spec)
set.seed(123)
wflw<- workflow() %>%
add_model(model_spec) %>%
add_recipe(recipe_spec)
error<-tryCatch({
set.seed(123)
wflw_fit<-fit(wflw,df_train)
error<-0
}, warning = function(w) {
print(w)
return(0)
}, error = function(e) {
print(e)
return(1)
})
if(error>0){
msg<-paste('Loss-Function failed...',title)
}else{
print('kein Error')
}
}
infoTest <- cvResamples$splits[[1]] %>% analysis()
infoTest %>% glimpse()
infoTrain <- cvResamples$splits[[1]] %>% assessment()
infoTrain %>% glimpse()
myPlot<-cvResamples %>%
tk_time_series_cv_plan() %>%
plot_time_series_cv_plan(
date, value,
.facet_ncol = 1,
.interactive = TRUE
)
View(cvResamples)
View(infoTrain)
View(infoTest)
#print(myPlot)
stop('Finished!')
Shuffled except date-var:

holdout seems to play a role...
model_spec <- exp_smoothing(
error = 'additive'
,trend = 'none'
,season = 'none'
# ,damping = 1
# ,smooth_level = 1
# ,smooth_trend = param6
# ,smooth_seasonal = param7
) %>%
set_engine("smooth_es",holdout=TRUE,silent=FALSE)
model_spec<-parsnip::eval_args(model_spec)smooth::es() (via modeltime wrapper) Noticed 5: msg: <simpleWarning: The exogenous variables contain NAs! This may lead to problems during estimation and in forecasting. Substituting them with 0.>
... is preventing the plotting in smooth (via modeltime)... code add...
colVect <- c('value','xReg')
data <- data %>% dplyr::mutate_at(all_of(colVect),as.numeric)
data <- data %>% dplyr::mutate(xReg=lag(xReg,n=1))
# comment this out to get the effect...
data <- data %>% dplyr::mutate_at(vars(matches("xReg")), ~tidyr::replace_na(.x,mean(.x,na.rm=T)))Update: msg-issue: Solved, must be an internal from modeltime/tidymodels, because plotting of extracted model works as expected. Even with NA-msg.
Plotting an extracted model via smooth:
wflw_fit<-fit(wflw,df_train)
myModel<-wflw_fit[['fit']][['fit']][['fit']][['models']][['model_1']]
myPlot<-plot(myModel,7)
print(myPlot)
stop()Doing the same (plotting extracted model) with ADAM brings:
Fehler in if (any(noVariability) && any(all.vars(formulaToUse) %in% names(noVariability))) { : Fehlender Wert, wo TRUE/FALSE nötig ist
myModel <- adam(df_train,"ANN", silent=FALSE,h=12, holdout=TRUE)
myPlot <- plot(myModel,7)
#myPlot<-ggplotly(myPlot)
print(myPlot)
stop()Can't reproduce it native.
I can reproduce the chart-design (native) by setting h=1
idx=120
y<-rnorm(idx, mean=15, sd=5)
x<-cbind(
x1 = rnorm(idx, mean=15, sd=5)
,x2 = rnorm(idx, mean=15, sd=5)
)
fitXregs <- data.frame(x=x,y=y,stringsAsFactors=F)
fitXregs <- fitXregs %>% dplyr::relocate(y)
myModel <- adam(fitXregs,"ANN",silent=FALSE,h=1,holdout=TRUE)
I' m not sure if this is right- comparing to the book-chapter....

https://openforecast.org/adam/SES.html
Hinting with ggplot+plotly
idx=120
y<-rnorm(idx, mean=15, sd=5)
x<-cbind(
x1 = rnorm(idx, mean=15, sd=5)
,x2 = rnorm(idx, mean=15, sd=5)
)
fitXregs <- data.frame(x=x,y=y,stringsAsFactors=F)
fitXregs <- fitXregs %>% dplyr::relocate(y)
myModel <- adam(fitXregs,"ANN",silent=TRUE,h=10,holdout=FALSE)
#myModel <- smooth::es(df_train,model='ANN',h=1,holdout=TRUE,silent=TRUE)
y <-myModel[['data']][['y']]
myFitted <-myModel[['fitted']]
myResiduals <-myModel[['residuals']]
myFc <-myModel[['forecast']]
myFuture <- data.frame(forecast=myFc)
myData <- data.frame(actual=y,fitted=myFitted,residuals=myResiduals)
dataLen <- nrow(myData)
myData <- dplyr::bind_rows(myData,myFuture)
myData <- myData %>% dplyr::mutate(id=row_number()) %>% relocate(id)
myIntercept <- myData[dataLen,'id']
# wide to long
library(reshape2)
subToLong = myData[,c(1,2,3,5)]
myData = melt(subToLong, id=c("id"))
myPlot <- ggplot(myData)
myPlot <- myPlot + geom_line(aes(x=id, y=value, colour=variable))
myPlot <- myPlot + geom_vline(xintercept=myIntercept, linetype="dashed", color = "#FF6900",size = 0.5)
myPlot <- myPlot + scale_colour_manual(values=c("black","red","blue"))
myPlot <- ggplotly(myPlot)
print(myPlot) 
Noticed 6: It also has a problem with: 'h=1'. Since modeltime had the same for a while, it seems to be a common problem in fc-plots. Anyway, this hint was fun :-).

Finally: fixing the hint... finding the lost one step forward fc (h=1, ~conditional mean)... :-).

[...]
dataLen1 <- nrow(myData)
hintPoint <- myData[dataLen1,c('id','value')]
myPlot <- ggplot(myData)
myPlot <- myPlot + geom_line(aes(x=id, y=value, colour=variable))
myPlot <- myPlot + geom_vline(xintercept=myIntercept, linetype="dashed", color = "#FF6900",size = 0.5)
myPlot <- myPlot + scale_colour_manual(values=c("black","red","blue"))
myPlot <- myPlot + ggtitle(title)
myPlot <- myPlot + geom_point(data=hintPoint,aes(x=id, y=value), colour="green",size=3)
myPlot <- ggplotly(myPlot)
print(myPlot)Are you using the most recent version of the smooth package? I cannot reproduce this. Here is what I get for the first part of your code:

Also please make sure that you use the latest version of the greybox package
I noticed yesterday, that my local versions are way to old. I am working on multiple instances, sorry.
Windows 7, greybox 1.0.5.41001, smooth 3.1.6.41004, R version 4.0.5 I get the following:
idx=120
y<-rnorm(idx, mean=15, sd=5)
x<-cbind(
x1 = rnorm(idx, mean=15, sd=5)
,x2 = rnorm(idx, mean=15, sd=5)
)
fitXregs <- data.frame(x=x,y=y,stringsAsFactors=F)
fitXregs <- fitXregs %>% dplyr::relocate(y)
myModel <- adam(fitXregs,"ANN",silent=FALSE,h=10,holdout=TRUE)
idx=120
y<-rnorm(idx, mean=15, sd=5)
x<-cbind(
x1 = rnorm(idx, mean=15, sd=5)
,x2 = rnorm(idx, mean=15, sd=5)
)
fitXregs <- data.frame(x=x,y=y,stringsAsFactors=F)
fitXregs <- fitXregs %>% dplyr::relocate(y)
myModel <- adam(fitXregs,"ANN",silent=FALSE,h=1,holdout=TRUE)
The same code can be simplified to (for reproducibility purposes, so that I do not need to load tidyverse packages):
idx <- 120
x<-cbind(
y = rnorm(idx, mean=15, sd=5),
x1 = rnorm(idx, mean=15, sd=5),
x2 = rnorm(idx, mean=15, sd=5)
)
fitXregs <- as.data.frame(x, stringsAsFactors=F)
myModel <- adam(fitXregs,"ANN",silent=FALSE,h=10,holdout=TRUE)But this is what I get:

And this is with h=1:

So, I still cannot reproduce this.
I assume that this can be closed now.
Should be environment related.
Hello, I get different model-graphs with the same data and model-definitions in adam, auto_adam and smooth::es() when setting silent=FALSE. Is this normal, and what do the x-axes of the plots represent?
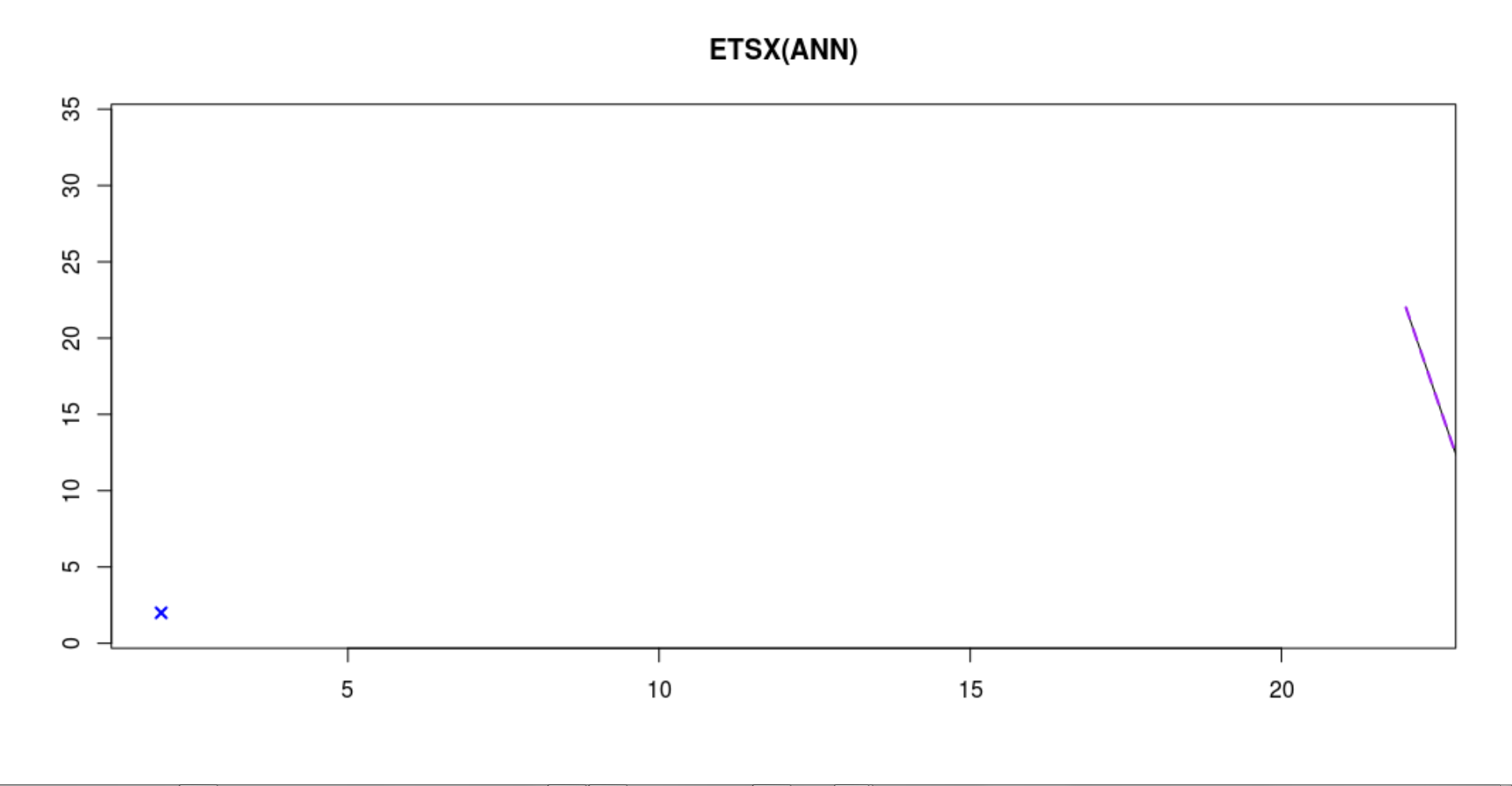
I also get this graph fitting ADAM, while cross validation for loss-function. What does the x on the left side mean?