Thanks @JosieLikesCats for flagging these at such an early stage, they are indeed very interesting sequences.
I'm sorry, I can't really provide much insights yet, but I would very much appreciate if you could try to answer some questions that come to mind: Do you happen to have raw reads for these - this would be very helpful to validate that the sequences are real and not for example due to coinfection. Are there other possible explanations for these? Do you know how closely related the individuals are who were sequenced?
Where in Limpopo and Gauteng are these from? Google maps says ~400km but could the Limpopo ones also be from a place closer to Gauteng province?
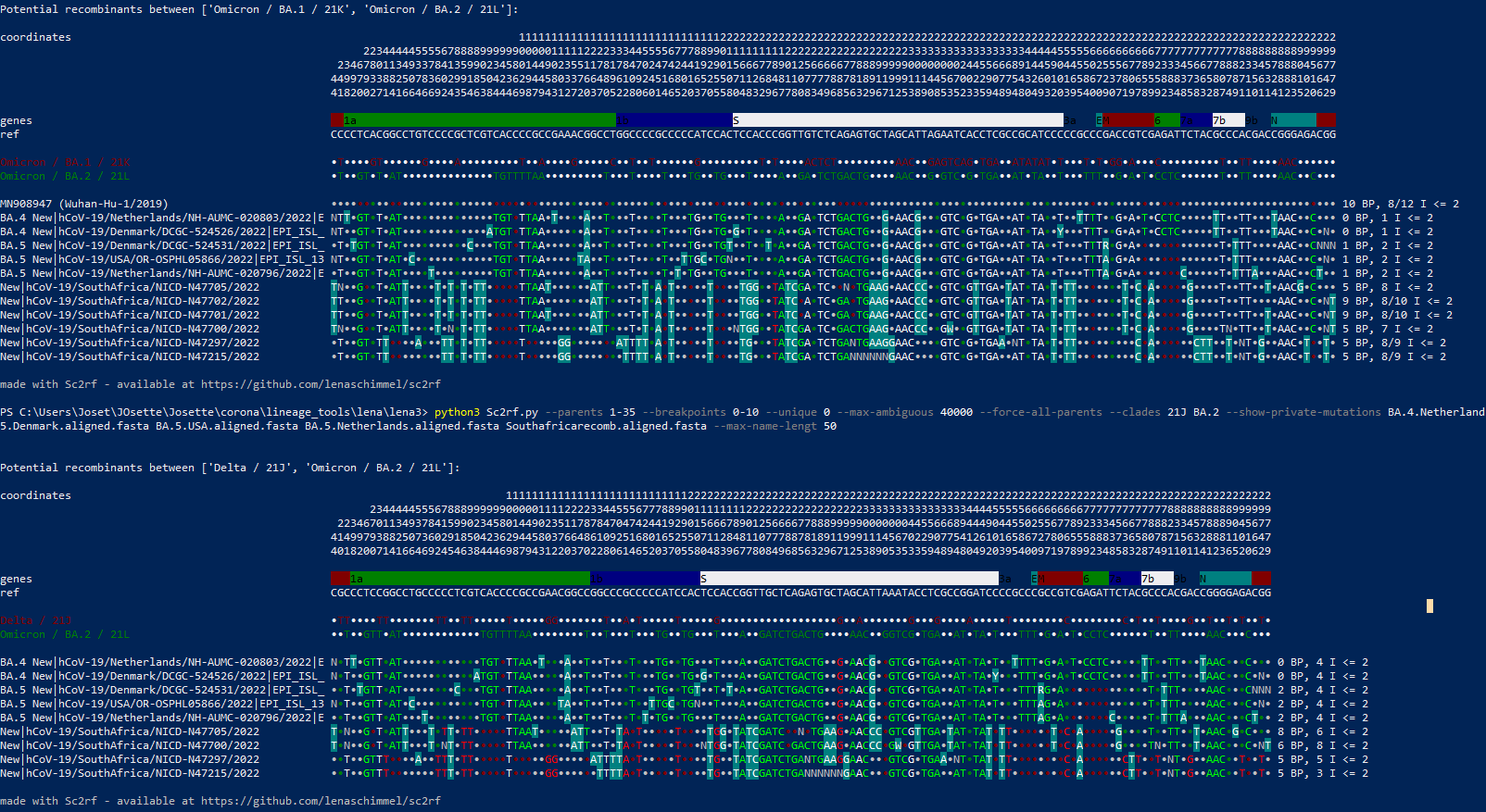
These sequences should be run through sc2rf to check for recombination - first impression is that it's very recombinanty. But also that both sequences seem to have some things in common. Very speculatively, these could represent different results of intra host recombination - but one needs to look at this in more detail.
It may be worth splitting this issue up and separate the proposals but for now to keep things simple I think it's fine to keep them both together.
Here are screenshots from Nextclade to save everyone some time:








Hi everyone, I'm just opening this issue to highlight that there are several sequences with an unusual mutation pattern in our most recent upload from South Africa, which will potentially represent two new lineages if more sequences are detected. The teams in our genomic surveillance network (NGS-SA) as well as our public health institute (NICD) are closely monitoring the sequences and cases in the country. These new constellations have been detected only in a small proportion of recent data, and our cases remain low.
I know these do not yet meet requirements for designation, as there are only N=4 and N=3 (2 available on GISAID, last 1 will be released tomorrow) sequences for each constellation, but we thought they would probably be of interest and picked up here/on Twitter eventually. For now, please see below for some details and the major mutation profiles for the two groups of sequences.
N=4 constellation 1 Earliest sequence: 28 June 2022 Most recent sequence: 29 June 2022 Circulating: Gauteng, South Africa Nextclade assigns 21M but flags lots of private mutations (mainly 21J), pango assigns Unassigned/B.1.1.529
Genomes EPI_ISL_13830378 EPI_ISL_13830377 EPI_ISL_13830376 EPI_ISL_13830375
N=3 constellation 2 Earliest sequence: 13 June 2022 Most recent sequence: 24 June 2022 Circulating: Limpopo, South Africa Nextclade assigns 21J but flags lots of private mutations (mainly 21K/21L), pango assigns XD
Genomes EPI_ISL_13830379 EPI_ISL_13830380
Evidence constellation1_defining_aa_changes.xlsx constellation2_defining_aa_changes.xlsx
Spike mutations in constellation 1 only, relative to Omicron: R21G, F486P, P621S, A706V Spike mutations in constellation 2 only, relative to Omicron: S477D Shared mutations relative to Omicron BA.4/5: L18F, T19R, W152L, E156del, F157del, R158G, F186L, G446D, T1117I Notably both clusters have a second silent nt change in L452R not present in BA.4/5. There are some significant differences outside spike (see attached mutation profiles). The sites 213, 371, 373, 375, 376, 408, and 764 are not reliably covered by the data, so they cannot be confirmed yet. UShER tree (including 7th sequence to be uploaded): https://nextstrain.org/fetch/genome.ucsc.edu/trash/ct/singleSubtreeAuspice_genome_1ef08_5f410.json (in a previous Usher tree they clustered near XD).