@emmamendelsohn is this similar to the outbreak history layer @noam started in that immunity is inferred from previous outbreak case data? Or is the immunity layer a separate seroprevalence dataset as suggested in #79? If based on case counts this will be included in PR #76 which contains daily outbreak histories in both the short and long term. In these histories, once an outbreak occurs the impact has an exponential decline, both spatially and as time progresses.
As an aside, SpatRasters produced with terra::writeRaster(..., gdal=c("COMPRESS=LZW")) were nearly half the size as when saving long-form data as parquet using compression = "gzip", compression_level = 5. Both .parquet and .tif versions are in the data/outbreak_history_dataset/ folder.
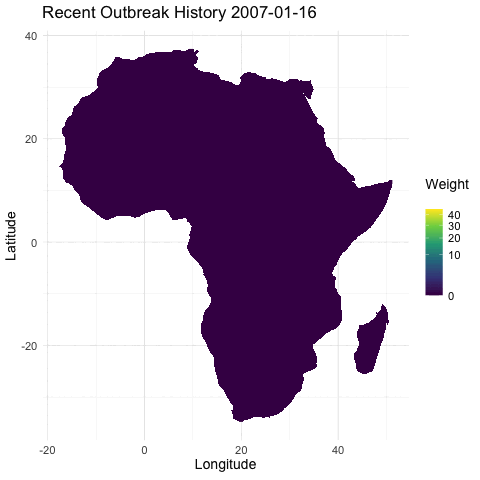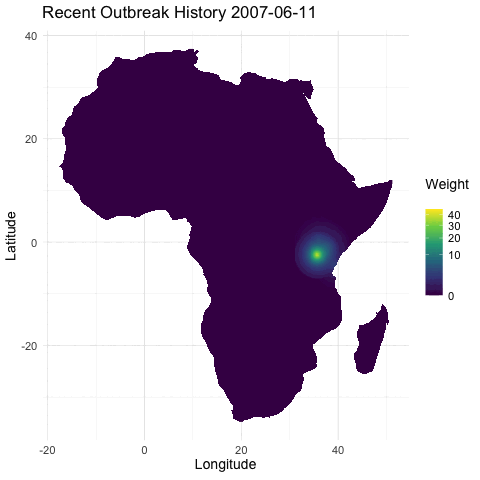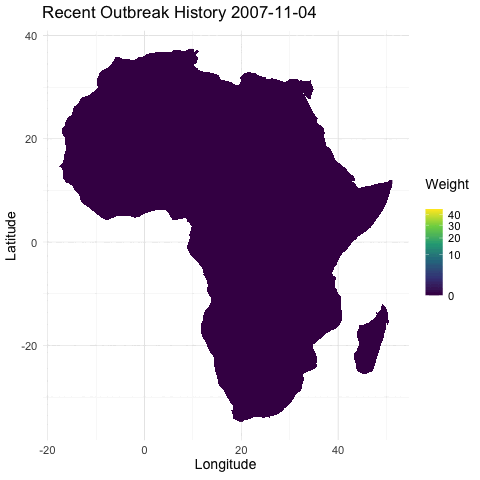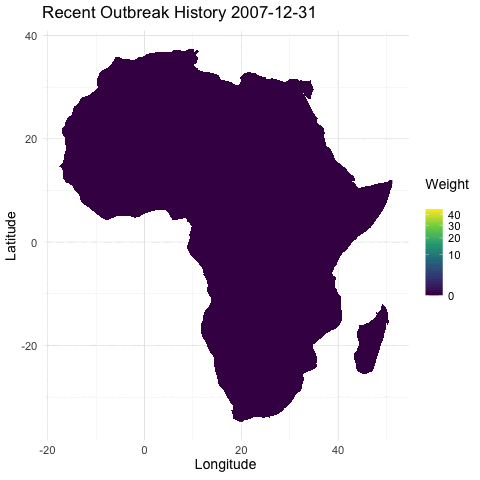
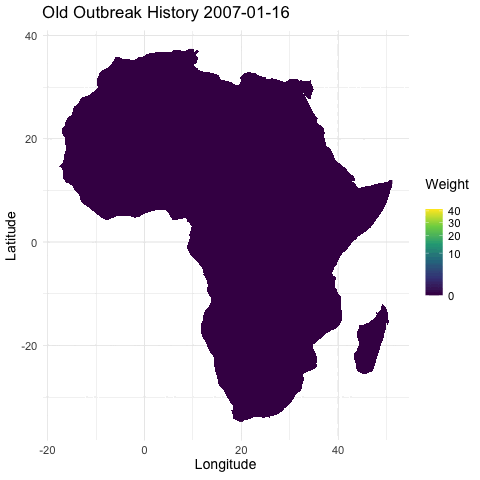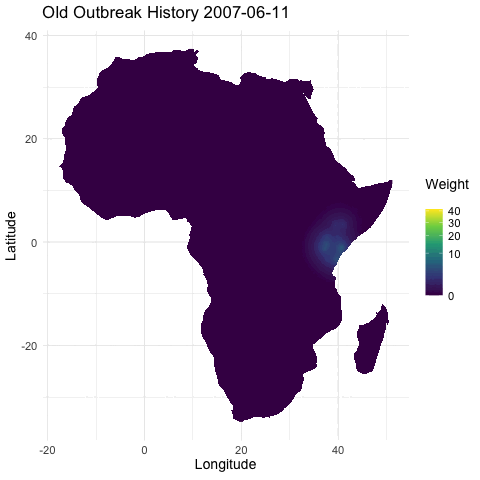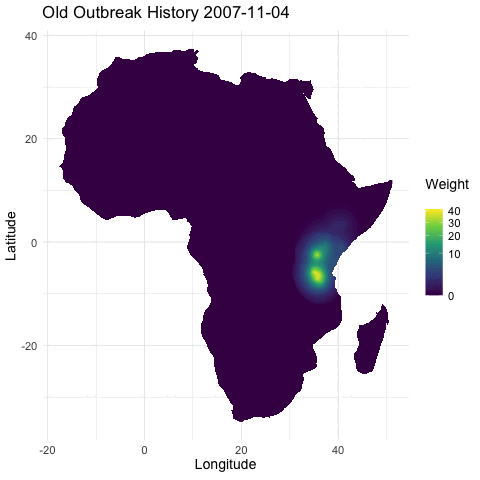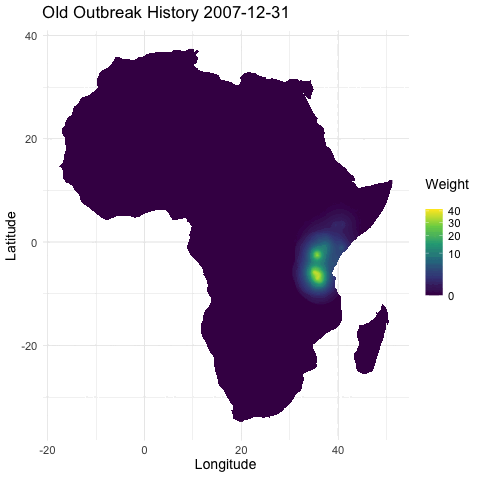
[ ] Separately, create a layer for whether there has been an outbreak in the region within the last year, as a first pass for capturing spread. (In a future iteration this could be something like distance to nearest outbreak in the last year.)
When averaging to the ADM level: