Hi,
It is a wonderful project and I am personally trying to make the reward function that you mentioned in your paper using Self-Organising Maps (SOMs).
For this I used both Morgan Fingerprint and Maccs Fingerprint as features. But the clusters in SOM are not converging and are scattered in
I also tried with features like Molecular weight, Synthetic accessibility, LogP, number of rings, QED score and Natural Product likeness. But still it did not converge.
So, I wanted to ask, what features did you use for creating these SOMs. I just want to create Kinase SOM and Specific DDR1 SOM.
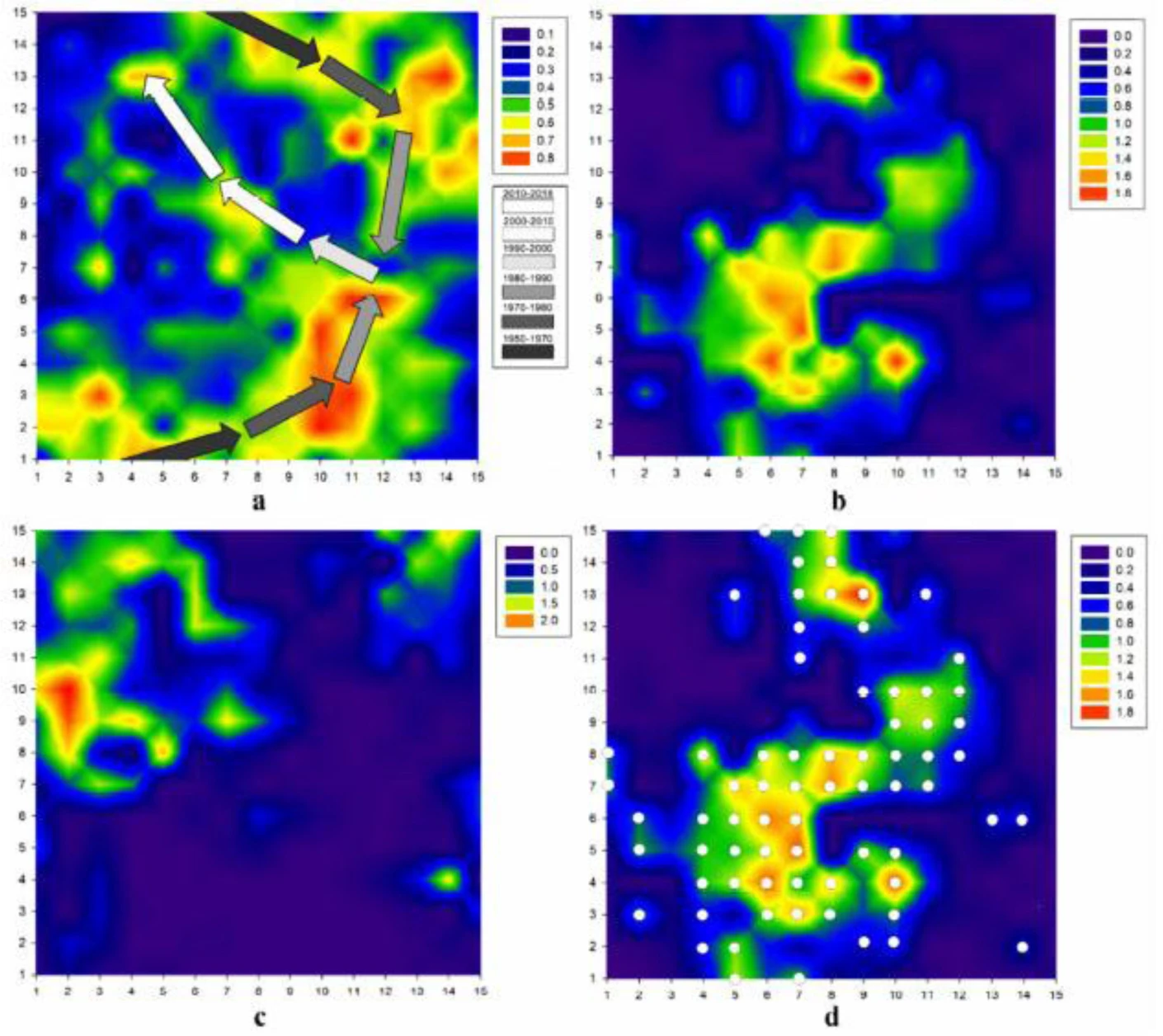
The (b) and (c) in this picture:
Hi, It is a wonderful project and I am personally trying to make the reward function that you mentioned in your paper using Self-Organising Maps (SOMs).
For this I used both Morgan Fingerprint and Maccs Fingerprint as features. But the clusters in SOM are not converging and are scattered in
I also tried with features like Molecular weight, Synthetic accessibility, LogP, number of rings, QED score and Natural Product likeness. But still it did not converge.
So, I wanted to ask, what features did you use for creating these SOMs. I just want to create Kinase SOM and Specific DDR1 SOM. The (b) and (c) in this picture: