Hi Lu Lu,
Just to elaborate, I have plotted the train loss that is contributed to by the pde with the test loss below.

The weird thing is that although the same metric (the pde) is used to evaluate these two losses, and the training and test points are located in the same region, the test loss increases as the training loss decreases.


 Example of points sampled: Boundary points: 400 Domain points: 1500 Test points: 1000
Example of points sampled: Boundary points: 400 Domain points: 1500 Test points: 1000 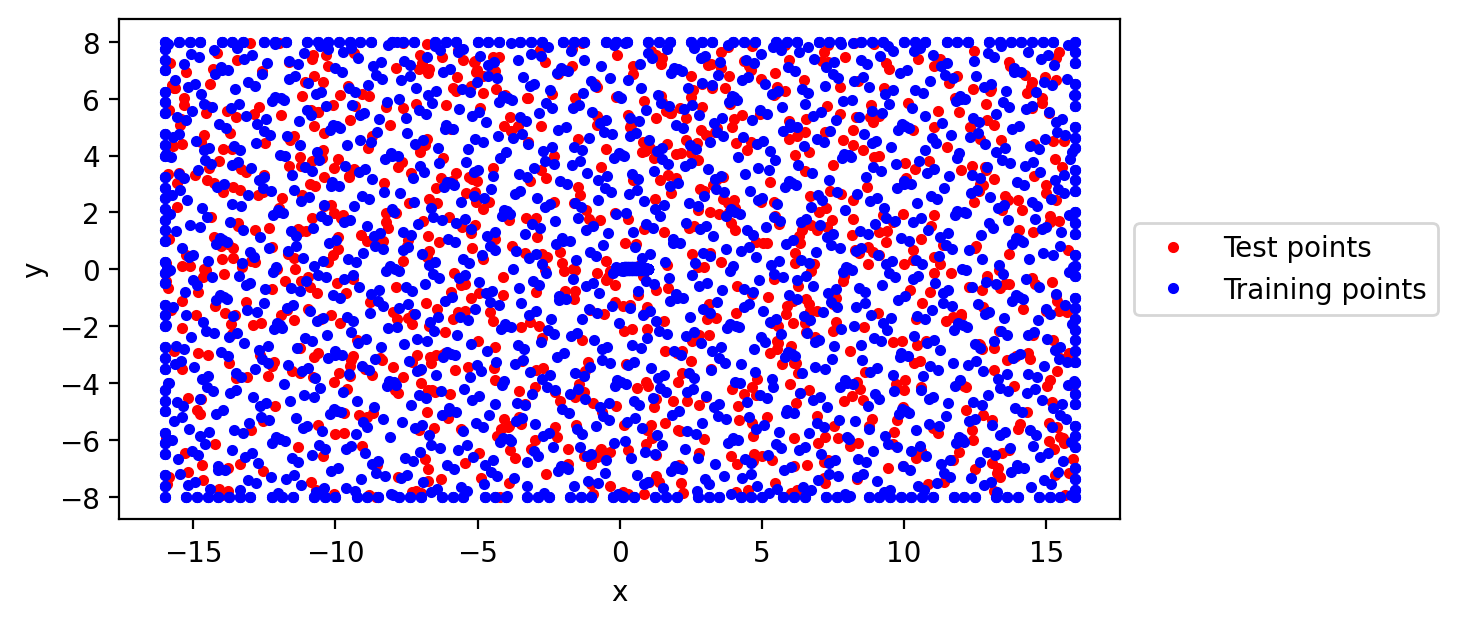
Hi @lululxvi,
I have been getting a problem where my test loss is opposite from my train loss (as my training loss decreases, test loss increases proportionally)
Example of the loss plot: Example of points sampled:
Boundary points: 400
Domain points: 1500
Test points: 1000
Example of points sampled:
Boundary points: 400
Domain points: 1500
Test points: 1000

PDE used:
Do you have any suggestions that I can try out?
Thank you very much.