I suspect the order of your images from the mrc_list.txt input, from this line:
ls -d -1 $dis/* >> $DATADIR/mrc_list.txt
Does not match the order of the CTF parameters parsed from cryosparc_P11_J4_003_particles.cs. The .cs file parameters are in the same order as the .star file from the EMPIAR 10028 entry.
Dear all,
Thanks for sharing the awesome work! I have been exploring with the latest version of CryoDRGN2 recently. I was trying to perform homogeneous reconstruction using
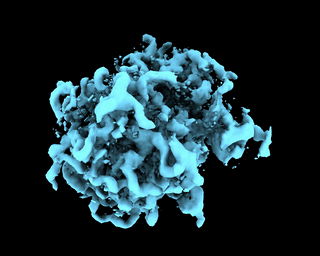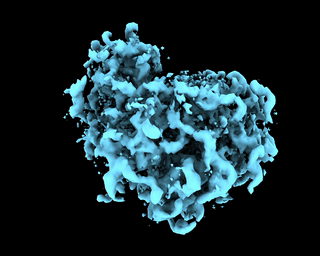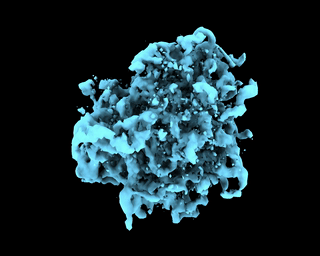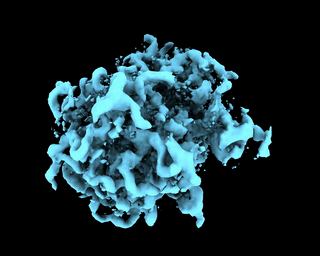
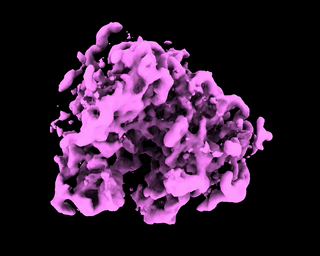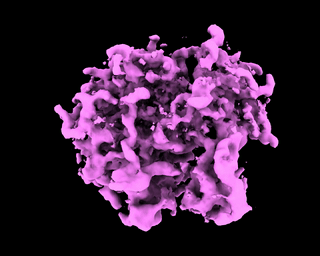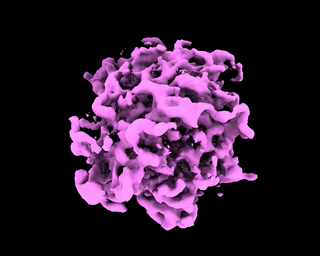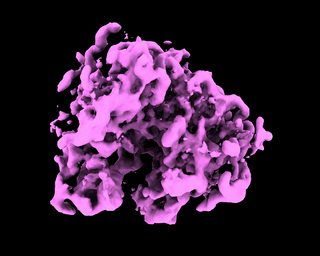
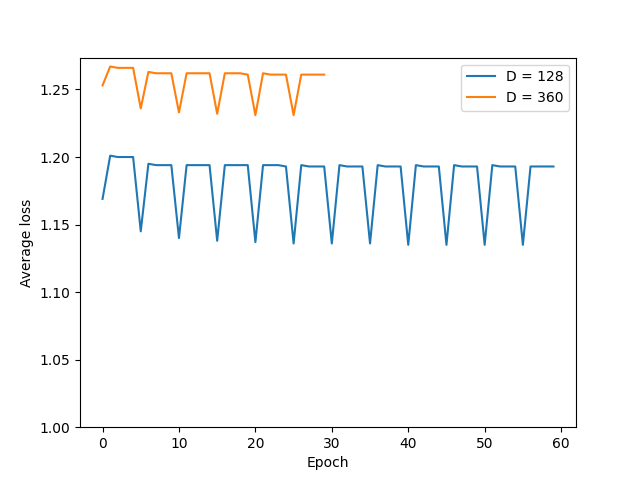
abinit_homofor EMPIAR-10028. However, after plotting out the loss it appears that the training did not converge after 60 epochs:The periodic pattern in the loss seems to coincide with the pose search update of 5 epochs reported in the paper, but I'm 100% certain if the periodic loss is attributed to the pose search steps. The resulted final volume of homogeneous reconstruction is below:
Steps to reproduce
ctf.pklfrom here. (script I used)cryodrgn downsample $DATADIR/mrc_list.txt -D 256 \ --max-threads 64 \ -o $DATADIR/particles.256.mrcs \ --chunk 50000
cryodrgn downsample $DATADIR/particles.256.txt -D 128 \ --max-threads 64 \ -o $DATADIR/particles.128.mrcs