thanks for your submission @DomBennett - in Repository: AntonelliLab/outsider can you give the url instead please
Closed DomBennett closed 4 years ago
thanks for your submission @DomBennett - in Repository: AntonelliLab/outsider can you give the url instead please
Done!
we're discussing now
thanks for your submission. @annakrystalli has been assigned as handling editor
Hello @DomBennett!
Just starting with the editors checks now. Will get back to you with any initial feedback and next steps shortly.
Many thanks again @DomBennett for your submission. A really interesting package!
I've completed the editor checks and am happy to start looking for reviewers. In the meantime, there are a couple of monior issues the initial checks have thrown up.
oldrelThere's a test failure that pops up in examples when running on oldrel. Forking your repo and adding:
language: r
r:
- oldrel
- releaseto travis.yml to also test on the old release of R indicates there's an API change between 3.4 and 3.5 R that affects package functionality, as oldrel fails.
In particular:
Checks
Throw an error in one of the module_details examples:
Running examples in ‘outsider-Ex.R’ failed
The error most likely occurred in:
> base::assign(".ptime", proc.time(), pos = "CheckExEnv")
> ### Name: module_details
> ### Title: Look up details on module(s)
> ### Aliases: module_details
>
> ### ** Examples
>
> library(outsider)
> # return table of ALL available modules on GitHub
> # NOT RUN - takes too long
> # (available_modules <- module_search())
>
> # look-up whether specific module exists
> # NOT RUN
> # repo <- 'dombennett/om..goodbye.world'
> # (suppressWarnings(module_exists(repo = repo)))
> repo <- 'dombennett/om..hello.world'
> (module_exists(repo = repo))
dombennett/om..hello.world
TRUE
> # get details on specific module(s)
> (module_details(repo = repo))
Error in if (inherits(X[[j]], "data.frame") && ncol(xj) > 1L) X[[j]] <- as.matrix(X[[j]]) :
missing value where TRUE/FALSE needed
Calls: module_details -> lapply -> FUN -> as.matrix.data.frame
Execution haltedTests
Again the same function fails the tests
* checking for unstated dependencies in ‘tests’ ... OK
* checking tests ...
Running ‘test-all.R’ [10s/74s]
ERROR
Running the tests in ‘tests/test-all.R’ failed.
Last 13 lines of output:
Container status 'This is a mock'
────────────────────────────────────────────────────────────────────────────────────────
── 1. Error: module_details() works (@test-search.R#94) ───────────────────────────────
missing value where TRUE/FALSE needed
1: module_details(repo = c(repo, repo)) at testthat/test-search.R:94
2: lapply(X = github_res, FUN = as.matrix)
3: FUN(X[[i]], ...)
4: as.matrix.data.frame(X[[i]], ...)
I've checked the CI section in our guide and admittedly we don't have any guidance on this currently, so we are discussing updating it as a result. In the meantime, can I ask that you:
travis.yml to also test on devel, release and oldrel (I can make a PR if you like)?oldrel? If you need any help debugging, just let us know.Firstly, goodpractice() throws a couple of warnings. it seems a few files don't end in a new line. I'm not sure how consequential that is to functionality. In any case, it probably means the files aren't being parsed correctly during goodpractice checks so it's best to address it we can check all the code.
Preparing: lintr
incomplete final line found on '/Users/Anna/Documents/workflows/rOpenSci/editorials/outsider/R/identities.R'incomplete final line found on '/Users/Anna/Documents/workflows/rOpenSci/editorials/outsider/tests/test-all.R'incomplete final line found on '/Users/Anna/Documents/workflows/rOpenSci/editorials/outsider/tests/test-all.R'incomplete final line found on '/Users/Anna/Documents/workflows/rOpenSci/editorials/outsider/tests/testthat/test-outsider.R'incomplete final line found on '/Users/Anna/Documents/workflows/rOpenSci/editorials/outsider/tests/testthat/test-outsider.R'incomplete final line found on '/Users/Anna/Documents/workflows/rOpenSci/editorials/outsider/tests/testthat/test-test.R'incomplete final line found on '/Users/Anna/Documents/workflows/rOpenSci/editorials/outsider/tests/testthat/test-test.R'Finally an issue that needs addressing
require()
It is good practice to
✖ avoid the library() and require() functions, they change the
global search path. If you need to use other packages, import them. If you
need to load them explicitly, then consider loadNamespace() instead, or as
a last resort, declare them as 'Depends' dependencies.
R/test.R:18:22
Could you also please:
[](https://github.com/ropensci/onboarding/issues/282)If you could try and address these before review begins officially, that would be great. I'll ping back in when I've secured reviewers. In the meantime, feel free to reach out with any questions
Reviewers: Due date:
Hi @annakrystalli
Thanks for the editor checks. I hope I've managed to correct the issues you raised!
Do you have any advice on running tests across different R versions (oldrel, release and devel) locally? Looks like I'd need to set up multiple virtual environments and install and run tests in each one; a bit of a palava.
Thanks! Dom
Hi @DomBennett,
Thanks for the swift action!
So we've opened an issue in the dev_guide regarding this (feel free to contribute to the discussion) and have outlined a couple of suggestions.
The main approach to test locally would be to use Rocker Docker images with different tags. Get different version with eg docker run -v $(pwd):/pkg rocker/verse:devel Rscript "devtools::check('/pkg')".
In your case It will be a bit trickier, because the package needs access to Docker itself. But it's doable. If you write a dockerfile that installs the docker client on top of a Rocker image, and mounts the docker socket when running the container, it should work.
Hope this helps and let us know if you have any more questions 😸
Thanks @annakrystalli! I'll look into using rocker and keep an eye on the issue in the dev_guide.
We now have reviewers! 🎉 Thanks for agreeing to review @hrbrmstr & @nuest. Please feel free to reach out with any questions
Reviewers: @hrbrmstr, @nuest Due date: 2018-03-17
Please check off boxes as applicable, and elaborate in comments below. Your review is not limited to these topics, as described in the reviewer guide
The package includes all the following forms of documentation:
URL, BugReports and Maintainer (which may be autogenerated via Authors@R).For packages co-submitting to JOSS
- [x] The package has an obvious research application according to JOSS's definition
The package contains a
paper.mdmatching JOSS's requirements with:
- [x] A short summary describing the high-level functionality of the software
- [x] Authors: A list of authors with their affiliations
- [ ] A statement of need clearly stating problems the software is designed to solve and its target audience.
- [ ] References: with DOIs for all those that have one (e.g. papers, datasets, software).
Estimated hours spent reviewing: 6
Wow - quite a big project you've created there. I'm not sure I can cover everything in this first round of review, so please understand I'm putting in a few questions in the following comments.
Please find the full output of my review tests at https://nuest.github.io/outsider-review/index.nb.html
[Disclaimer: I did not do yet write my own module because of time constraints.]
0.0.0 seems curious to me, I suggest to either go with the .9000 versioning as described here if you think this is a development state, if you think you have a first working version (as the "active" repo status suggests), why not go with 0.1.0?paper.md should go into the base directory..-Notation is misleading. You normally don't want users of your package (be they normal or power users) to rely on such "internal" functions. What about using different prefixes?, e.g. moduledev_.. for developer functions?om.. notation really only for discovery on GitHub? If so, have you considered using GitHub tags instead? If you don't require specific names, people could enhance their existing R packages to become otutsider modules.DESCRIPTION metadata vs. om.yml: Can you elaborate on the reasoning to put the Docker user name in the DESCRIPTION into a custom field, and not into the om.yml ? At first sight it might be easier for users to have the DESCRIPTION file as standard as possible and put your own metadata in one place. Or the other way around: Why not put program and details into the DESCRIPTION file?module_list(repo = 'dombennett/om..mafft') is feasible to implement.README.md: I recommend to have an empty line after headlines and a space after the #/## in the source code for readabilitysysreqs issue, and re-check https://ropensci.github.io/dev_guide/building.html#package-dependencies.module_validate(..) to validate the formal structure of a module - I now see there is .module_check :-) - I suggest to put this "TODO" into a GitHub issue so that potential collaborators see this is an open task, and not have a function that a user cann access with ::: without any functionality. Or at least add a - _Is there a way to list all function names in a module?_ The Get Started vignette says "Once a function name is known of a particular .." - can I only do that via the help file? MAybe amodule_list(repo = 'dombennett/om..mafft')` is feasible to implement..warning("not implemented")`test())pkgdetails_get I personally would rely on the package desc to do the parsing - you already have a number of imports and this lightweight one should not hurt.module_travis - your approach means that the function will fail if the repo mooves, or if there is no internet connection... for me it's always a signal to reconsider the approach if you have a constant like a URL within a function body. I think it would be better to manage the "template" file within the codebase of the package.container.R: your package code probable precedes stevedore, but if you find yourself writing code around calling docker with sys::exec.. then it might be worth the effort to switch; things like running containers remotely (if that is what you mean by "remote hosting") would come very cheaply then;search.R:
tkn <- NULL to make it more obvious when requests fail because of a missing token (and not because the token is not valid etc.)info[['url']] <- paste0('https://github.com/', info[['repo']]) > reuse gh_url from the top of the same filemodule_exists is not implemented fully, see suggestion above (warning and/or GitHub issue)test-search.R, the "main" function testing in test-outsider.R, test-log.R (i.e. run actual workflows, capture the output, and see if the user is (not) shown what she should see). You could also run integration tests with incomplete modules and see if they fail at the expected point.test-docker.R
docker_pull, should be skipped on CRAN, see ?testthat::skip_on_cran, or you add your own test function skip_if_docker_is_not_available() (later I saw a comment "TODO: add skips if no docker" - go for it :-) ! )img variable from the previous test - that can quickly break. I suggest to either duplicate img, or move it outside of the test_that(..) functiondocker_img_rm() returns FALSEexpect_warning(expect_equal(attr(system2("docker", args = c("inspect", img), stdout = TRUE, stderr = TRUE), "status"), 1)), or with the package stevedore (also applies to test-container.R)test-console.R
expect_output(..), e.g. expect_output(outsider:::cat_line('cat this'), "cat this") (and also for celebrate() and comfort())outsider:::vars_get() - the function is only for testing, right? Then I think it would be better to add it to the test codebase, e.g. in test-all.RphylotaR and ape to Suggests because you use them in vignettesoutsider covers a pretty complex problem! With the existing examples of one very basic "hello world" and one (to me very) complex analysis pipeline, I see room for a further example that shows a bit more but is not that domain specific, e.g. loading some data from a CSV and running a regression model, saving that to a file. For me as a user it would be great to know how can I load my data, how call my function, and how access the output data.input_sequences.fasta does not work - would it be possible to let a user download (with R code) such a file and actually run that? I was a bit disappointed when I went though all previous steps and then was "stopped" unexpectedly.log_set already in the get started documentation."library(outsider)" statements in each section?module_help(repo = 'dombennett/om..mafft', fname = 'mafft') more helpful? Right now it only says "Run mafft". I think for the "story" of the vignette it would be much better to actually have some documentation there, e.g. reference papers and describe the function.FROM instruction in the Docker docs, WORKDIR, ..)
stevedore package, which allows them to do the buidling of Docker images from R directly.Please excuse my brevity at some points, and don't be discouraged at all but the length of the comments. I think the package is a useful approach to packaging workflows and tools in a user friendly way and a great usage of container technology.
It would be great to see this beeing picked up in your lab/your community of practice and see how people interact with each other's work based on outsider modules.
Hi @nuest,
Thanks for the super helpful review! Lots of new things and ideas to work on and with (... wish I had heard of stevedore beforehand!)
Many thanks indeed for your detailed review @nuest! And enjoy your upcoming holiday 😊
mine shall be completed today
Great. Thanks for the update @hrbrmstr
NOTE: I deliberately didn't read @nuest's review before going through my own package review. I then read @nuest's most excellent/thorough review and removed [most of, hopefully] any duplicate findings unless I felt the need to reinforce some.
Please check off boxes as applicable, and elaborate in comments below. Your review is not limited to these topics, as described in the reviewer guide
The package includes all the following forms of documentation:
URL, BugReports and Maintainer (which may be autogenerated via Authors@R).For packages co-submitting to JOSS
- [X] The package has an obvious research application according to JOSS's definition
The package contains a
paper.mdmatching JOSS's requirements with:
- [X] A short summary describing the high-level functionality of the software
- [X] Authors: A list of authors with their affiliations
- [X] A statement of need clearly stating problems the software is designed to solve and its target audience.
- [ ] References: with DOIs for all those that have one (e.g. papers, datasets, software).
Estimated hours spent reviewing: 5
paper.md IMO does contain a sufficient "statement of need" even though it's all in the Summarydevtools::test() and devtools::run_examples() all ran fine on macOS 10.14.4 RC / R 3.5.3stevedore. This is not a show stopper but it would reduce package size and compelexity and perhaps be less confusing to some R folks.(again, trying to not duplicate @nuest, but apologies if there are)
First: Bravo! This was pretty darned cool once I got it going.
Second: Definitely check out "whalebrew" (https://github.com/whalebrew/whalebrew) as I think that might be a good project to side-reference somewhere in the outsider docs (since it accomplished a similar but not duplicative goal) and also consider how they've structured their concept of "modules" (package): https://github.com/whalebrew/whalebrew-packages. In fact, it was really only when I "AHA!"'d the fact that outsider was "whalebrew for R" that I finally grok'd the package.
I shall now stop with the numerical itemizations :-)
I will now suggest something that will likely cause you to hurl something to me if we ever meet at an R conference: consider splitting the package into two separate packages, with the main outsider package being for the general, average, "will likely never build a Docker image in their lives but could make excellent use of existing outsider modules" R users with very targeted documentation and functionality and, perhaps "outsider.devtools" that focuses on making it as easy as possible for folks to build, test, and deploy outsider modules. I say that for a few reasons. First, the function list in the help index is extremely daunting for a large swatch of R users and most of the functions are for module builders. With a more focused, general-user-level outsider package you'd likely get more adoption and could focus all the documentation on just using modules. Further, I found the:
outsider problem) => process a bit…unsatisfying.
Speaking of which, to get to ^^ I actually built a module!! : https://github.com/hrbrmstr/om..nmap : but I'm not sure why the one error shows up on module_details() since I followed the directions to create the skeleton and push stuff up to the proper locations:
library(outsider)
repo <- "hrbrmstr/om..nmap"
module_search()
module_uninstall(repo)
module_install(repo)
module_details(repo)
## Error in paste(lines, collapse = "\n") :
## cannot coerce type 'closure' to vector of type 'character'
module_help(repo)
nmap <- outsider::module_import("nmap", "hrbrmstr/om..nmap")
x <- nmap("-v", "-oX", "/working_dir/outputfile.xml", "-sS", "a.b.c.d)
str(x)
xml2::read_xml(x$outputfile.xml)
## {xml_document}
## <nmaprun scanner="nmap" args="nmap -v -oX /working_dir/outputfile.xml -sS a.b.c.d" start="1553359840" startstr="Sat Mar 23 16:50:40 2019" version="7.70" xmloutputversion="1.04">
## [1] <scaninfo type="syn" protocol="tcp" numservices="1000" services=" ...
## [2] <verbose level="1"/>
## [3] <debugging level="0"/>
## [4] <taskbegin task="Ping Scan" time="1553359840"/>
## [5] <taskend task="Ping Scan" time="1553359841" extrainfo="1 total ho ...
## [6] <taskbegin task="Parallel DNS resolution of 1 host." time="155335 ...
## [7] <taskend task="Parallel DNS resolution of 1 host." time="15533598 ...
## [8] <taskbegin task="SYN Stealth Scan" time="1553359841"/>
## [9] <taskend task="SYN Stealth Scan" time="1553359842" extrainfo="100 ...
## [10] <host starttime="1553359840" endtime="1553359842">\n <status sta ...
## [11] <runstats>\n <finished time="1553359842" timestr="Sat Mar 23 16: ...If you do look at https://github.com/hrbrmstr/om..nmap you'll see that the markdown generated by the package generates invalid R code (no double quotes around the repo param values).
All of the recently above is likely summarised by "I think module building documentation and tooling could use a little more work, especially bits about getting data back from them" (between man pages and looking at an example of yours I obviously figured it out, but just before I did I almost would have preferred using whalebrew's nmap image and system2()). Once I finally worked through making a module I can completely see the utility of this package and think it's a great addition to the R universe.
Regardless of whether it is split into two packages, I would like to see an on-load/attach pre-flight check and/or pre-flight check function(s). For "users" this could check for Docker presence/version and any other items that are essential to the successful operation of the package. For "builders" it could also check for essential/recommended environment variables and/or anything else that you know might make it easier to do work with the package.
I think the functions themselves are properly documented for their functionality and the code style is fine, but I'm curious as to the inconsistent use of stringr functions. I'm also generally not a fan of adding a stringr dep vs a stringi dep, especially if you aren't using any "in stringr-only functions" since stringr depends on stringi and stringi is already a bear of a dependency on linux systems. In fact, I only see two FQ uses of it so I somewhat question the need for it at all. I didn't do a full check for whether it comes along for the ride with the Imports, but I don't think it does. So I'd suggest validating that and either moving to stringi or just using base R string functions (like you do with sub()/gsub()).
One bit of tension I also had as a module developer was the check "green" travis checks causing a warning message to be displayed. I think a further check could be made to ensure that the package is even being tested at all and then message() that vs warning() it.
Finally, as noted a few times, this package installed w/o a hitch as did using modules. That is most, most, most impressive for massive amount of complexity this package hides from the user. Very well done!
As someone who only puts his stuff on GitHub now to make it easier for all the R folks who are being led astray there by RStudio, (w/apologies to the team) rOpenSci, DataCamp, et al to find his stuff I'm kinda put off by the absolute need for GitHub. I 100% understand that supporting different social coding sites adds complexity but it also breeds diversity and avoids single points of failure. So, please consider not having a hard-dep on GitHub for module discovery.
What might make ^^ easier to deal with (though it requires a re-jiggering of module discovery and expectations) and to make it far easier for users in general is to mimic what whalebrew does (again, https://github.com/whalebrew/whalebrew-packages) and require modules to be under a single repo. You could do this via submodules so the code stays with the owner (like Hugo themes does) but is "registered" under there. Then, you could keep this single repo mirrored across GitLab, GitHub and other free and less harmful-to-humanity sites.
Speaking of modules… I'm already not a fan security-wise of the existing way GitHub is used to brazenly load Windows binary DLLs onto user systmems from GitHub and this goes a step further and puts the entire infrastructure of a containerized OS onto a local system pretty opqauely (yes, so does whalebrew). Great for abstraction, not great for system or analysis integrity. The winbin dance is going away soon and I think I'd feel a bit better about outsider modules if the "single repo with subdirs" approach was taken and there were hard-and-fast set of guidelines in place to be accepted into this new "outsider verified registry". Under the umbrella of rOpenSci these could be established in parallel with traditional onboarding practices but I'd rly argue that:
Unlike apt get / brew install Docker is the "Wild, Wild West" and GitHub is "Max Max". With outsider coming under the umbrella of rOpenSci I rly think the need for a more formal module registry is needed since — while it would not be rOpenSci's fault if something bad happened — an integrity or security incident could negatively impact rOpenSci's reputation. Unlike stevedore (where it's just a Docker API wrapper) this is very R specific and is also suggesting that this outsider approach is sound/safe for scientific computing (and I'm not saying that it can't be). So, to me, that kinda means there should be at least some type of badge/star/etc that goes next to "official/reviewd/clean/supported/safe/secure" outsider modules and ones that are just "out there". Perhaps I'm overthinking this and just paranoid (which is a part of the job description). So I'd def like all folks to weigh in here. I will also gladly craft a malicious module if it comes down to that :-)
I'm going to re-visit the package Sunday from scratch with fresh eyes to make sure I didn't miss anything not covered by @annakrystalli or @nuest.
Overall: amazing job; very useful concept and package and the execution is solid as-is tech-wise.
Thanks for the equally thorough review @hrbrmstr! 😃💯
Over to you @DomBennett. I think the reviews have provided really valuable feedback so look forward to hearing what you think.
As always, please feel free to ask for clarification or start a conversation on anything you'd like more input on. 👍
Thanks @hrbrmstr for the super thorough, well-thought and detailed review! I've got myself lots of things to be thinking about and getting on with.
Thanks @annakrystalli, I'll get to addressing these issues soon as.
Merci à toutes et tous
Just been lead to this via twitter - if you are interested in using stevedore but hit any hurdles @DomBennett, please feel free to contact me on the rOpenSci slack
Hello @DomBennett 👋
Any chance we could get an update on progress with responses to the reviews? Feel free to reach out if you are stuck on anything.
Hi @annakrystalli
I've still not started on any responses to the reviewers in earnest just yet. Currently juggling multiple things, and my hand-eye coordination is not the best.
Will try and get stuck in next week and send you a sit-rep along with any help requests I may have.
Thanks!
Hej allihopa!
Once again, thanks for the super-duper detailed and insightful reviews to the package. Lots of good advice and angles I hadn’t considered. Sorry for my lateness getting to this; I was juggling many things, plus I always have trouble approaching reviews for the first time. 😬
Given the complexity of this package and the high-level suggestions provided by the reviewers, I figured I should aim to be more communicative than in my previous submissions to ROpenSci where I just implemented the changes and dumped a great big message in the review. Here, I think a bit more to and fro is required.
So to start off with, I will outline below both the major and minor changes I intend to implement as well as the changes I would like a bit more discussion/help with before I implement anything. As I implement changes, I will then provide sit-reps on progress and setbacks.
outsider.devtoolsAs both reviewers indicated, the mixing of dev and user functions is confusing. I actually have no issue in creating a separate package and siphoning off the strictly dev related tools to this new package. I actually like this idea and think it is the cleanest approach. I assume ROpenSci would have no quibble in hosting the additional package.
Issue: https://github.com/AntonelliLab/outsider/issues/3
stevedoreI will definitely switch code to stevedore for outsider.devtools. This makes a lots of sense to reduce community-level code duplication and to support future features and docker dev within R. I am, however, a bit tentative about implementing stevedore for outsider-proper because of the potential issues for Windows users (https://github.com/richfitz/stevedore#windows-support). (@richfitz, would Windows users be unable to pull and run docker images without the need to install all the dependencies that you list?)
Issue: https://github.com/AntonelliLab/outsider/issues/4
@hrbrmstr, agreed. Other options should be available. It should be possible to create tools that can either search GitLab, BitBucket or whatever else. As an absolute failsafe, it should be possible to install a module from a local directory.
Issue: https://github.com/AntonelliLab/outsider/issues/5
Implement AppVeyor CI. This is a major issue for me as I’ve never used Appveyor! If I have any issues, I’ll let you, @nuest, know. I’m just going to see others do it and mimic.
Issue: https://github.com/AntonelliLab/outsider/issues/6
@nuest, I actually went for the mocks because I was having big issues with the time the tests were taking and I was having issues with uninstalling and reinstalling packages, as part of the module tests, on Travis. There were issues related to lib paths that I tried to fix using withr::with_temp_libpaths but only had mixed success.
I can solve the time issue by using skips. I also like testing for functional and non-functional modules. For these larger tests of interaction, I may encounter issues with the libpaths again. I’ll will try and fix these issues.
Issue: https://github.com/AntonelliLab/outsider/issues/7
I will splice the documentation between outsider and outsider.devtools. I will also try and create a middle of the road vignette that is not technical (biological sequence alignment) nor too simple (hello world). The advanced vignette will move to outsider.devtools and be enriched with all the stevedore goodies with extra links to pre-existing documentation.
Issue: https://github.com/AntonelliLab/outsider/issues/8
I have a few ideas for the future direction of outsider, many for which I have no idea how to implement. @nuest, I will do as you say and create them as separate issues.
Issue: https://github.com/AntonelliLab/outsider/issues/9
@nuest suggests I drop the need for “om..”. I can do this. I actually already make use of the GitHub description in addition to the “om..” and the presence of om.yml. I think I did these three tests initially because I thought the more tests there are for determining whether a repo was a module or not the better. In hindsight, this is probably not necessary. It may be better to just use the GitHub labelling system. A label like “outsider-module” should work just fine. (But, then, it will be a different system on other code-sharing sites.)
@hrbrmstr suggests I add all “approved” modules to a single repo and that I might be able to make use of GitHub’s submodules feature. I looked into submodules early on in outsider’s development and found that they were not universally praised by all those who used them. More simply, I could just have a list of approved repo URLs in the main outsider repo. (This might allow for non-GitHub modules as well.)
Ultimately, however, I didn’t want to create lots of work for myself nor others – such as those at ROpenSci who would, presumably, be hosts for this “approved” modules listing. I understand the need to ensure that modules are not malicious, but I do worry that trying to police them may be a task too great to take on. Additionally, are we even able to determine whether a module is safe or not? I might be able to determine the non-malicious status of a module in my own field of research (biodiversity, macroevolution, phylogenetics) because I’d likely know the people and recognise the software being run. But I’m not sure if I could outside of my field; we researchers tend to write quite obfuscated code by default. The worst-case scenario might then be a module is approved, which then turns out to be malicious. Would I be held responsible for any potential fallout? Also, do we have any suggestions on what sorts of malicious things could be achieved through an outsider module? And, are there anyways of ensuring they’re not doing these sorts of things?
My thinking to save community-effort is to allow users to download any module they like from whichever code-sharing site but to provide warnings along the lines of:
Beware: outsider modules have the potential of introducing malicious software.
This repo has # many watchers and # many stars. The maintainer of the repo is #.
Please make sure you are confident this is a legitimate outsider module.
Note, outsider maintainers take no responsibility for code developed by external developers.
Are you sure you wish to proceed?
(Y/N)Perhaps this warning could appear on any code that has too few stars/watchers?
(Presumably I’m being paranoid now considering that whalebrew has an approved list. Plus this all assumes that outsider gets picked up.)
Issue: https://github.com/AntonelliLab/outsider/issues/10
paper.md, om.yml, allowing: dropping om.. requirement, removal of metadata from DESCRIPTION.module_check/validatedesc.module_travis: don’t make dependent on URLsearch.R issuesexpect_output (I didn’t know this existed!), move strict testing functions to test-all.RstringrIssue: https://github.com/AntonelliLab/outsider/issues/11
@annakrystalli, there’s a ROpenSci slack? Would it be possible to have access to it?
Thanks!
Geia sou @DomBennett! 😜
Many thanks for your detailed response and apologies for the delay in mine!
I see the invitation to the slack has gone through so welcome! Feel free to ask questions for broader feedback there.
Re:
As both reviewers indicated, the mixing of dev and user functions is confusing. I actually have no issue in creating a separate package and siphoning off the strictly dev related tools to this new package. I actually like this idea and think it is the cleanest approach. I assume ROpenSci would have no quibble in hosting the additional package.
Yes that should be fine, it would be deemed in scope. We'll need to check that:
If all these checks pass, the package should also be considered reviewed.
Re: Your security concerns, I like the approach you've proposed. Let's wait to hear reviewers' opinions.
Let me know if you need any more info from me. Looking forward to @nuest & @hrbrmstr input!
Hello again @hrbrmstr & @nuest. Was wondering if you had any feedback for @DomBennett regarding the major changes he is proposing?
Hallå hallå,
Quick update on progress
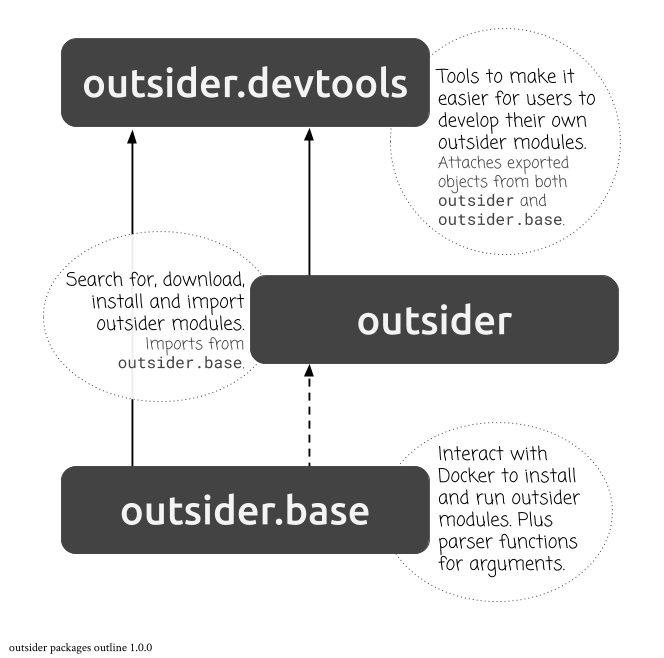
I’ve actually decided to split the package three ways: outsider.base, outsider and outsider.devtools.
Why? I actually found it quite hard to determine what should be exported objects if I just had outsider and outsider.devtools. But now there’s quite a clear line of dependence and clear statements of purpose of each package.
Other updates
outsider and outsider.base, along with all the previous modules I made, are now functional (as tested so far, at least).outsider can install modules from GitHub, GitLab, Bitbucket, URL or downloaded file path.Still TODO
outsider.devtoolsHi all, just catching up on the many good plans. Looks very promising and a huge step forward for the package(s). Here are some answers/comments:
outsider get's so much attention that the first security issue arises! Be aware, educate users, but don't fix problems that don't exist.om..+in:name from you GH search string :-).outsider.base: since outsider.devtools devtools will always rely on outsider (right?), I think factoring out the common base was not a strict requirement. But small R packages are getting more common, so :+1: Comments on code changes
(I browsed through the code changes only.)
tests/testthat/test-download.R: testing url_make(), why do you not test the content of the URL but only the type?remotes to install from the different sources! Have you considered supporting remotes::install_git(..) as well? Many research institutes run their own GitLab instances these days and a plain git link should always work then. If you want to save some work, maybe add to the documentation that the regular URL also works for any third party/self-hosted git collaboration site. test-github.R)I applaud your efforts in getting things really right, but have to admit given the amount of things to do I am a bit overwhelmed and have a hard time giving feedback. I think you're on the right track! If you need some specific feedback, please feel free to mention me in GitHub issues, and if I don't get back to you within a couple of days, ping me via email or Gitter.
Thanks @nuest -- I will act on your early feedback here!
On a general point to the reviewers, please don't feel overwhelmed or obliged to respond to all the points I raise in this thread. I just want to lay out my thinking and show the evolution of the package(s) in an open and accessible way.
Thank you for the feedback @nuest! ✨ And please don't worry @DomBennett.
I can somewhat understand where @nuest is coming from because the splitting up of the package does add additional checking that perhaps wasn't envisaged at the start. For completing the review, I would refrain from tackling "future dev ideas" and try and limit the "A healthy plethora of other things" on your TODO list to changes that are strictly necessary for functionality or are responding directly to reviewers comments. (Although by all means, track them as issues in the repositories themselves). As I'm sure you know, development can and always does, continue beyond review, in response to a broader user base (which may even lend a hand too!).
But just to dispell any worries you or @nuest have and given that @hrbrmstr might not be available to continue, I will be happy to do the final checks to help spread the load a bit. So carry on @DomBennett and just let us know when you think you're ready for final checks. 👍
Ey up 👋 (Yorkshire speak for hey there)
Just dropping by to see how it's going. 🙂
As always, feel free to reach out with questions / request for feedback if you're stuck (although I appreciate it's more than likely a finding the time thing!)
Hi @DomBennett, just checking in on this review. Hope everything is going well your end. Let me know if you need feedback on anything.
Hi @annakrystalli -- sorry! Just been really slow on this: lots of other things plus summer. Will keep it turning over with updates as I go along.
No problem! Thanks for the update and yeah, just keep us in the loop of developments. 🙂
Enjoy 🏖☀️!
Hi all!
Sorry for my massively slow updates: lots of things were happening, this project is way bigger than any of my other ROpenSci projects.
ssh featureoutsider keeps all of the outsider.base functions hidden – avoiding confusion. For the developer, they need to learn the exported outsider.base functions, upon calling the outsider.devtools these functions are made public.stevedore has not been used, desc has not been used. These packages appear useful but since I’ve already coded the base without them I see no reason in switching to using them just not. If new features are required or errors can be fixed with these packages, they can always be added at a later stage without impacting the user experience.sysreqsdb. It should be possible for linux machines but would require extra scripts, not directly available via any packaging suites (apt, RPM). Plus then there is the question of docker flavour (ce or ee?). I know too little.I hope these changes are going in the right direction. Let me know of any feedback or issues you have.
Thanks!
Hey @DomBennett ,
This is great work!
@nuest or @hrbrmstr do you perhaps have time to comment on some of the changes made? I'm happy to do the final checks on all three packages when all the minor fixes still pending are addressed, but your higher-level input at this point on the decisions being made could be really useful.
@DomBennett , should we decide on the minor tasks that need completing to be able to draw the line when they are done? Are you happy to complete the checklist you've got in your issue or are there some that you consider enhancements and therefore not necessarily critical to the issues raised by reviewers?
ssh featureOnce we establish what's left to complete, it would be good to get an idea of a time frame in which they might be complete. 👍
Sound good?
Hi @annakrystalli !
Thanks for your response. I've been quietly beavering away at the list. I have addressed all of the issues now with one or two cavets:
(Additionally, there are come Appveyor issues to do with installation of packages for the devel R release. And there seems to be an issue with installation of fs on Travis for my oldrel.)
How should we proceed next?
Thanks!
Great work @DomBennett !!
Well done for chipping away. It sounds like you are in a place where we could potentially start wrapping the review up! All 3 packages need to be looked at though. As I will be doing it, I'm planning to take next weekend to spend some time on it.
I'll also have a look at and enquire about the issues you are having with testing on devel and oldrel. In the meantime, I wonder if this might help: https://github.com/r-lib/fs/issues/90
Thanks @annakrystalli !
In terms of the CI issues, I tried fixing the failing Travis build for oldrel for the outsider package based on that issue you directed me to. I tried installing fs by specifying it as a binary package, this prevented the fs error but then I ended up with a new installation error for a different package. So I figured this is clearly a general compilation issue on travis... which led me to this issue: https://github.com/wesm/feather/issues/154
I followed the travis YAML guidance specified in the above issue and found that .... it didn't fix the problem, same old installation error with fs.
So for now, I've dropped oldrel from the outsider/.travis.yml. I can try it out again later. I figure at some point the error would be fixed regardless of my efforts.
One puzzle: why should this be an issue for outsider but not outsider.devtools? They have virtually the same .travis.ymls (Or at least, outsider.devtools has all the same dependencies.)
Please check off boxes as applicable, and elaborate in comments below. Your review is not limited to these topics, as described in the reviewer guide
The package includes all the following forms of documentation:
URL, BugReports and Maintainer (which may be autogenerated via Authors@R).For packages co-submitting to JOSS
- [x] The package has an obvious research application according to JOSS's definition
The package contains a
paper.mdmatching JOSS's requirements with:
- [x] A short summary describing the high-level functionality of the software
- [x] Authors: A list of authors with their affiliations
- [x] A statement of need clearly stating problems the software is designed to solve and its target audience.
- [x] References: with DOIs for all those that have one (e.g. papers, datasets, software).
Estimated hours spent reviewing: 5
Thanks for your patience with the review. I've completed the review for outsider. Will get back to you with the review on the other two packages later today.
Firstly well done on the great work you've done on the package. It feels like the splits you decided on work well and it seems thhe functionality works well. There are still however a few issues to work through, not suprising given the extensive refactoring you've done. So let's work through them:
Running
devtools::install()on source code returns the following error
Running /Library/Frameworks/R.framework/Resources/bin/R CMD \
INSTALL \
/var/folders/8p/87cqdx2s34vfvcgh04l6z72w0000gn/T//RtmptOYBoM/outsider_0.1.0.tar.gz \
--install-tests
* installing to library ‘/Library/Frameworks/R.framework/Versions/3.6/Resources/library’
ERROR: dependency ‘outsider.base’ is not available for package ‘outsider’
* removing ‘/Library/Frameworks/R.framework/Versions/3.6/Resources/library/outsider’outsider.base package remote needs referencing in the DESCRIPTION file like so:
Remotes:
AntonelliLab/outsider.baseoutsider.baseHowever, there is a problem with installation of outsider.base from GitHub still, with this error:
Downloading GitHub repo AntonelliLab/outsider.base@master
'/usr/bin/git' clone --depth 1 --no-hardlinks --recurse-submodules https://github.com/DomBennett/om..hello.world /var/folders/8p/87cqdx2s34vfvcgh04l6z72w0000gn/T//RtmpZ8wDI8/remotes14452732bf805/AntonelliLab-outsider.base-b8708e3/om..hello.world
dyld: Library not loaded: @rpath/libswiftCore.dylib
Referenced from: /Applications/Xcode.app/Contents/Developer/usr/bin/xcodebuild
Reason: image not found
git: error: unable to locate xcodebuild, please make sure the path to the Xcode folder is set correctly!
git: error: You can set the path to the Xcode folder using /usr/bin/xcode-select -switch
Error: Failed to install 'outsider.base' from GitHub:
Command failed (71)
In addition: Warning message:
In system(full, intern = TRUE, ignore.stderr = quiet) :
running command ''/usr/bin/git' clone --depth 1 --no-hardlinks --recurse-submodules https://github.com/DomBennett/om..hello.world /var/folders/8p/87cqdx2s34vfvcgh04l6z72w0000gn/T//RtmpZ8wDI8/remotes14452732bf805/AntonelliLab-outsider.base-b8708e3/om..hello.world' had status 71I solved it by running:
sudo xcode-select --switch /Library/Developer/CommandLineTools/So all good but, given the heavy system requirents, it might be good to consider collecting common troubleshooting advice in a vignette like you have in outsider.devtools docs.
check()There is a couple of test failures for me. It seems the gitlab api call is returning unauthorised.
CONTRIBUTIONG.md── Test failures ───────────────────────────────────── test-all ────
> library(testthat)
> test_check("outsider")
Loading required package: outsider
----------------
outsider v 0.1.0
----------------
- Security notice: be sure of which modules you install
── 1. Error: gitlab_repo_search() works (@test-gitlab.R#4) ────────
HTTP error 401.
Backtrace:
1. outsider:::gitlab_repo_search(repo = "DomBennett/om..hello.world")
2. jsonlite::fromJSON(search_url)
3. jsonlite:::parse_and_simplify(...)
4. jsonlite:::parseJSON(txt, bigint_as_char)
5. jsonlite:::parse_con(txt, bigint_as_char)
7. base::open.connection(con, "rb")
── 2. Error: gitlab_search() works (@test-gitlab.R#10) ────────────
HTTP error 401.
Backtrace:
1. outsider:::gitlab_search()
2. jsonlite::fromJSON(search_url)
3. jsonlite:::parse_and_simplify(...)
4. jsonlite:::parseJSON(txt, bigint_as_char)
5. jsonlite:::parse_con(txt, bigint_as_char)
7. base::open.connection(con, "rb")
══ testthat results ═══════════════════════════════════════════════
[ OK: 56 | SKIPPED: 0 | WARNINGS: 3 | FAILED: 2 ]
1. Error: gitlab_repo_search() works (@test-gitlab.R#4)
2. Error: gitlab_search() works (@test-gitlab.R#10)
Error: testthat unit tests failed
Execution halted
1 error ✖ | 0 warnings ✔ | 1 note ✖I also get a note:
Non-standard file/directory found at top level:
‘CONTRIBUTING.md’use_build_ignore("CONTRIBUTING.md") to add the file to .Rbuildignore.goodpractice()A couple of super minor points thrown up.
✖ avoid long code lines, it is bad for
readability. Also, many people prefer editor windows
that are about 80 characters wide. Try make your lines
shorter than 80 characters
R/gitlab.R:64:1
✖ not import packages as a whole, as this can
cause name clashes between the imported packages.
Instead, import only the specific functions you need.I'm tempted to say that because you're importing outsider.base that might be ok. In any case you could consider whether you indeed need the whole package imported? This might help with your consideration.
spell()Only one valid typo I could find! Awesome!
WORD FOUND IN
Analsis phylogenetic_pipeline.Rmd:104
alignment_example.gif on pkgdown site landing page missing.is_module_installed documentation confusing. It seems to be a copy of module_uninstall docs.Details entry in module_import reads: If program is successfully removed from your system, TRUE is returned else FALSE, is that right?.Get Started article is that users not hugely familiar with the command line might benefit from being walked though what each argument being passed is doing is mafft(arglist = c('--auto', 'input_sequences.fasta', '>', 'output_alignment.fasta'))outsider:::. You can leave them if you prefer but I would generally recommend focusing on extensive docs for user facing functions.CONTRIBUTING.md. See usethhis::use_code_of_conduct().om..* module docs.Module repos need a bit more documentation.
[ ] README need more detail. Someone arriving at the repo should be able to get atleast a high level understanding of what the repo contains. Perhaps you could develop a README template with basic info (perhaps even a function in outsider.devtools a bit like usethis::use_readme_rmd() to create them). You can that you customise for each module. with some brief info on what the module does and how to use an example function, including relevant code, much like any other package README.
[ ] It would be good in the module function documentation to either define the parameters or link to official documentation for the module/function and it's parameters. ALthough fname('--help') can be enough, sometimes it's not, eg. bamm('--help') does not yield much info.
[ ] Function descriptions also need a bit more info on what they do, eg in om..bamm, here should be a short description of what BAMM is. The details field (eg Bayesian Analysis of Macroevolutionary Mixtures) returned as part of module_details() would in most cases suffice, ideally with links to further information and references.
I won't hold up completing the review for this but I encourage you to both take the time to complete it for your modules and consider what best practice you want to model for users and how you can make it easy for them to follow said practice. This could have a large impact on usability of the modules produced by users.
I've completed most checks but want to spend a bit more time working with the package so back again tomorrow.
Thanks very much @annakrystalli! Super helpful.
I've got through and made the following changes:
outsider.devtoolsoutsider.base was designed to be used like this.is_module_installed, fixed!om.. docsI'm going to work on your suggesed changes to these next. Will work on them in conjunction with updating outsider.devtools.
Thanks!
Great work! And many apologies for dragging my heels with finishing the reviews. Will have it with you by the end of the weekend. Thanks for your patience!!
Sent from my iPhone
On 7 Jan 2020, at 15:57, Dom Bennett notifications@github.com wrote:
Thanks very much @annakrystalli! Super helpful.
I've got through and made the following changes:
Added a remotes section to DESCRIPTION Ditto for outsider.devtools Added Xcode and Rtools information to the "Getting started" page I added a skip to the GitLab API checks if no token is found. I've added information about access tokens to CONTRIBUTING and the documenation. CONTRIBUTING is now being ignored GP checks: No lines are too long now GP checks: I'm going to suggest ignoring the "do not import an entire package" because outsider.base was designed to be used like this. Spelling, fixed! Documentation: Alignment GIF, fixed! Documentation: GH link, added! Documentation: is_module_installed, fixed! Documentation: `module_imports, fixed! Documentation: more info on command-line, added! Documentation: more info on security, added! Documentation: I've removed "private functions" from the reference on the website Code of conduct, added! om.. docs
I'm going to work on your suggesed changes to these next. Will work on them in conjunction with updating outsider.devtools.
Thanks!
— You are receiving this because you were mentioned. Reply to this email directly, view it on GitHub, or unsubscribe.
Thanks! And no bother, I've been super slow myself.
Hi @annakrystalli!
As a quick update, I've updated the documentation for om..bamm plus a few other modules (om..astral, om..raxml, om..wikit, om..pathd8).
I've also created a template README.Rmd as part of the module skeleton for outsider.devtools, https://github.com/AntonelliLab/outsider.devtools/blob/master/inst/extdata/template_README.Rmd
The new template suggests developers provide a detailed example with output, information on some key arguments and links and references to the original program.
I've not had time to update the documentation for all the current modules that I maintain. But I've created an issue to remind that it needs done: Issue - Update documentation
Awesome work @DomBennett ! I really like where these packages are going. It's been long journey so well done for sticking with it!
OK so here's the review for outsider.devtools. Generally only minor points, primarily on the documentation. I also realise that because outsider.base is not outward facing, it does not need a formal review. So after responding to the outsider.devtools review points and a quick double check, I believe we will be good to go! 😃🙌
Please check off boxes as applicable, and elaborate in comments below. Your review is not limited to these topics, as described in the reviewer guide
The package includes all the following forms of documentation:
URL, BugReports and Maintainer (which may be autogenerated via Authors@R).For packages co-submitting to JOSS
- [ ] The package has an obvious research application according to JOSS's definition
The package contains a
paper.mdmatching JOSS's requirements with:
- [ ] A short summary describing the high-level functionality of the software
- [ ] Authors: A list of authors with their affiliations
- [ ] A statement of need clearly stating problems the software is designed to solve and its target audience.
- [ ] References: with DOIs for all those that have one (e.g. papers, datasets, software).
Estimated hours spent reviewing:
All good! :+1:
goodpractice::gp()── GP outsider.devtools ────────────────────────────────────────────
It is good practice to
✖ avoid long code lines, it is bad for
readability. Also, many people prefer editor windows
that are about 80 characters wide. Try make your lines
shorter than 80 characters
R/utils.R:87:1
──────────────────────────────────────────────────────────────────── memmory advanced.Rmd:136
repo's advanced.Rmd:192
structre intermediate.Rmd:191
Ts advanced.Rmd:211
webiste advanced.Rmd:197[ ] Again, the documentation for unexported function is confusing as functionality in the documentation doesn't work when you try and run it unless you know how to access internal functions.
[ ] No examples found for docker_build, docker_push, git_upload, module_identities, module_skeleton, module_travis, module_upload,
[ ] how do pkgdetails_get and description_get differ?
[ ] In docker_build help, what does value logical mean. ie what would it mean if the function returned TRUE?
docker.R?
outsider.devtools Coverage: 90.36%
R/docker.R: 56.25%module_skeleton using the same flpth argument, it ovewrites all files without warning. You might want to look at checking for this and asking users to confirm overwrite (eg have a look at function usethis::write_over()).DESCRIPTION file should have an Author field for a named person to formally take responsibility for the module. Using this would also allow users to create citations for their modules via citation().docker_build and other docker_cmd in both outsider.base.module_build with build_image = FALSE installs the module as a package in your local R library. In which situation would this be useful? It also means you don't need to build an image to use the functions in a module. You could be more explicit about what module building with and without building the image achieves. Perhaps you could point to and elaborate on these points in the FAQ section?Some of these suggestions are minor and just personal opinions so feel free to ignore if you don't agree.
[ ] Installation instructions in README should be changed as there is no need to manually install packages as you now have specified remotes.
[ ] (Super minor!) In the quick guide block of code, it would be more legible with blank lines in between lines of code, eg:
library(outsider.devtools)
# construct a skeleton file structure for the module
module_path <- module_skeleton(program_name = 'echo', flpth = getwd())
# check the file structure
module_check(flpth = module_path)
# look-up key identifying names: R package name, Docker image name
module_identities(flpth = module_path)
# build the R package and Docker image
module_build(flpth = module_path, tag = 'latest')
# test the module
module_test(flpth = module_path)[ ] Installation instructions in the Install section should be amended to reflect the automated installation of outsider and outsider.base.
[ ] .
These code snippets are called Docker images.
Not sure that's an accurate description of what a docker image is. My understanding is is that a docker image is an executable with an environment configured within which to run it. Overall and throughout documentation, I suggest you try to be precise with the terms you use in the docs and be consistent throughout. Docker is already somewhat confusing so misconceptions and inaccuracies in outsider docs could make it even more confusing. If unsure, have a look at the docker docs themselves.
[ ] Very minor point but visually, a list structure would be a lot more effective:
This process comes in a series of steps:
- generate core files and folders
- create Docker image
- document and build R package
- try the module
- upload to GitHub and Docker Hub
[ ] .
The above code will create an outsider module with the name om..echo
I feel it would be useful to point out here that the module is created at flpth.
[ ] I know it feels like repetition but nevertheless, I feel it would be useful to show the resulting structure of the module from running module_skeleton() using output of command line tree <temp-folder-path>/om..echo :
/var/folders/8p/87cqdx2s34vfvcgh04l6z72w0000gn/T//RtmptA6YLp/om..echo
├── DESCRIPTION
├── R
│ ├── functions.R
│ └── import.R
├── README.md
├── examples
│ └── example.R
└── inst
├── dockerfiles
│ └── latest
│ └── Dockerfile
└── om.yml
5 directories, 7 files[ ] The import and use of outsider.base functions in documentation or workflows is confusing because there is no function documentation in the outsider.devtools pkgdown reference section yet the package from which eg outsider_init() is imported from is not obvious to the user. I think it might be best to use the reexports approach (eg see how it looks in the docs and handled through roxygen in the R scripts in devtools)
[ ] .
Additionally, the developer would then not need to document all the arguments of the external program. For many external programs there may be hundreds of arguments, all of which are likely to be already documented;
This is a great approach if documentation exists for the external module. But it might be good to encourage folks to check this and if there is none as part of the program, to atleast linked to appropriate documentation in the module docs.
[ ] .
For more information, look up “object documentation in R”.
Best to provide the actual link to more information: http://r-pkgs.had.co.nz/man.html
[ ] .
The WORKDIR sets the starting directory when a command is passed to the Docker image.
This sentence feels confusing. It sets the working directory. As I commented above, try to be precise and consistent with tricky terminology and if unsure, have a look at the docker docs themselves. eg here's the intro to WORKDIR instruction docs.
The WORKDIR instruction sets the working directory for any RUN, CMD, ENTRYPOINT, COPY and ADD instructions that follow it in the Dockerfile
[ ] .
Using our Dockerfile that has been preloaded with the installed R package
Nothing is really preloaded in a dockerfile. It is just a text file. The computational environment is configured during the buiding of the docker image. I personally would rewrite this paragraph instead describing step by step what happens when the dockerfile builds.
[ ] .
To demonstrate the program’s complexity, here is the command’s signature
That's rather unusual terminology. Do you mean command's syntax? Again I suggest you keep to conventional terminology.
Thanks very much @annakrystalli !
I've fixed a few of your comments already will get to work on the others soon.
pkgdetails_get and description_getdocker_ functionsmodule_skeletondocker_build and other docker_cmd in both outsider.basebuild_image=FALSE in module_buildflpthoutsider.base import/exports. Attempt to clarify and normalise by mimicking well-known package structuresGreat to see the progress here. Please let me know if there are any specific things I should take a look at again, I'm afraid looking at everything is not an option the coming weeks.
@DomBennett :+1: for endurance :100:
Thanks @nuest !
@annakrystalli I've gone through my checklist and made all the requested changes (I hope!). Notable updates are:
module_skeleton, using usethis::write_overoutsider.devtools coverage improveddocker_build and docker_cmd exisiting in both the dev and base packages. They are both functionally the same, but in the dev version they print to console the bash commands to make it more transparent what is going on.build_image=FALSE. I do this in the example to separate the R building from the image building so that I don't need to explain both processes in the same section. One reason to avoid building an image is if you are updating the package code but haven't changed the Dockerfile. Would that be useful info to add to the FAQs?outsider.base, https://antonellilab.github.io/outsider.base/reference/index.html. I was hoping the devtools "reexports" page would automatically link to this documentation rather than the rddr.io documentation but I'm not sure how pkgdown links.Let me know if you need me to do anything else!
Approved! 🥳
Thank you @DomBennett for submitting (and going on this epic review and development journey!) and @nuest and @hrbrmstr for your reviews! 🙌
To-dos:
[](https://ropensci.org)"pkgdown website, fix its URL to point to https://docs.ropensci.org/package_name and deactivate the automatic deployment you might have set up, since it will not be built centrally like for all rOpenSci packages, see http://devdevguide.netlify.com/#docsropensci. In addition, in your DESCRIPTION file, include the docs link in the URL field alongside the link to the GitHub repository, e.g.: URL: https://docs.ropensci.org/foobar (website) https://github.com/ropensci/foobarDESCRIPTION via rodev::add_ro_desc().[](https://ci.appveyor.com/project/individualaccount/pkgname).For JOSS submission
This package has been reviewed by rOpenSci: https://LINK.TO/THE/REVIEW/ISSUE, @ropensci/editorsShould you want to acknowledge your reviewers in your package DESCRIPTION, you can do so by making them "rev"-type contributors in the Authors@R field (with their consent). More info on this here.
Welcome aboard! We'd love to host a blog post about your package - either a short introduction to it with one example or a longer post with some narrative about its development or something you learned, and an example of its use. If you are interested, review the instructions, and tag @stefaniebutland in your reply. She will get in touch about timing and can answer any questions.
We've put together an online book with our best practice and tips, this chapter starts the 3d section that's about guidance for after onboarding. Please tell us what could be improved, the corresponding repo is here.
Huzzah!
Thanks so much @annakrystalli!
Also big thanks to @nuest and @hrbrmstr!
Is it OK to add you all as "rev" in the R package DESCRIPTION?
Hey @DomBennett, thanks for transferring the repos over, I've returned full admin rights back to you.
Just double-checking with the editors team whether all will live in rOpenSci. In the meantime, could you briefly explain what the outsider-testsuite repo is/contains? Is it user facing?
Submitting Author: Dom Bennett (@dombennett)
Repository: https://github.com/AntonelliLab/outsider Version submitted: 0.0.0 Editor: @annakrystalli
Reviewer 1: @nuest
Reviewer 2: @hrbrmstr Archive: TBD
Version accepted: 0.1.0
Scope
Please indicate which category or categories from our package fit policies this package falls under: (Please check an appropriate box below. If you are unsure, we suggest you make a pre-submission inquiry.):
Explain how and why the package falls under these categories (briefly, 1-2 sentences):
The aim of the package is to enable R users to install and run, within R, software, packages and programs external to R. This allows users to more readily create workflows and pipelines that are portable, reproducible and make use of a wide range of available methods outside of R.
The target audience are data scientists, bioinformaticians and general scientists interested in running external programs within R.
There are many available packages that allow the interfacing between software external to R (e.g. packages for interacting with python or java). There does not, however, seem to be any packages that make use of docker (www.docker.com) to import external software as this package does. The closest packages to this one appear to be things like rocker. Rocker, however, uses docker to make R more portable, not external software accessible within R.
Technical checks
Confirm each of the following by checking the box. This package:
Publication options
JOSS Options
- [x] The package has an **obvious research application** according to [JOSS's definition](https://joss.readthedocs.io/en/latest/submitting.html#submission-requirements). - [x] The package contains a `paper.md` matching [JOSS's requirements](https://joss.readthedocs.io/en/latest/submitting.html#what-should-my-paper-contain) with a high-level description in the package root or in `inst/`. - [x] The package is deposited in a long-term repository with the DOI: - (*Do not submit your package separately to JOSS*)MEE Options
- [ ] The package is novel and will be of interest to the broad readership of the journal. - [ ] The manuscript describing the package is no longer than 3000 words. - [ ] You intend to archive the code for the package in a long-term repository which meets the requirements of the journal (see [MEE's Policy on Publishing Code](http://besjournals.onlinelibrary.wiley.com/hub/journal/10.1111/(ISSN)2041-210X/journal-resources/policy-on-publishing-code.html)) - (*Scope: Do consider MEE's [Aims and Scope](http://besjournals.onlinelibrary.wiley.com/hub/journal/10.1111/(ISSN)2041-210X/aims-and-scope/read-full-aims-and-scope.html) for your manuscript. We make no guarantee that your manuscript will be within MEE scope.*) - (*Although not required, we strongly recommend having a full manuscript prepared when you submit here.*) - (*Please do not submit your package separately to Methods in Ecology and Evolution*)Code of conduct