-
Hi,
I'm running compare_genomes pipeline and am having this error:
Error in file(file, "r") : cannot open the connection
Calls: read.nexus -> scan -> file
In addition: Warning message:
…
-
Hello, I found a problem. Because of how the file names work, it is possible to break the program with sufficiently large orthogroups
`sh: 1: cannot create BG_IM_KAI8734412.1-BG_IM_KAI8742320.1-BG_…
-
Hello, I have run Orthofinder successfully. However, I'm not sure if the SpeciesTree_rooted_node_labels.txt file is a tree file based on a Single_Copy_Orthologue ? Looking forward to your reply . Tha…
-
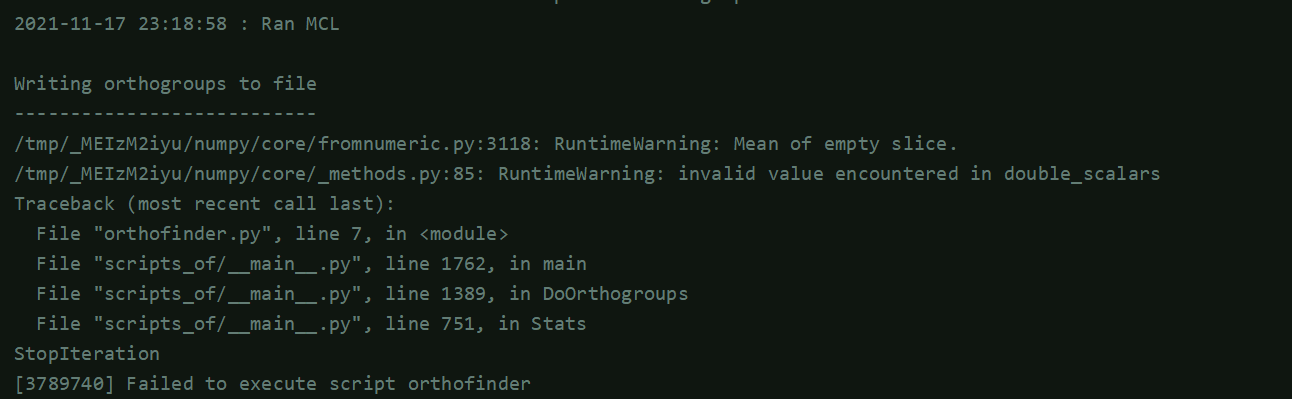
2021-11-17 23:18:58 : Written final scores for species 38 to graph file
2021-11-17 23:…
-
Hi, I have a dataset of 2905 genomes and would like to use Orthofinder to generate a phylogenetic tree. I used 72 cores and it has been 7 days and it is still running. Do you have an estimation of how…
ghost updated
2 years ago
-
Hi David,
Thank you for your previous reply. I would really appreciate your help if you could clarify my following doubts:
1. Can orthofinder produce single copy orthologue file using the nucleotide…
-
I've started playing around with this module. All in all, it works quite well. I've been using it to represent the major intersections of large (20) numbers of categories. Its quite easy to plot the t…
-
i have installed orthofinder by conda
orthofinder -hOrthoFinder version 2.4.0 Copyright (C) 2014 David Emms
SIMPLE USAGE:
Run full OrthoFinder analysis on FASTA format proteomes in
orthofind…
-
Hi David,
Recently,I ran 163 plant species in Orthofinder for one week and I got this error.
Reconciling gene trees and species tree
---------------------------------------
Outgroup: Porphyra_um…
-
Hello,
While working on the orthogroups PR, I ran into an issue with the way proteins are managed in GNB.
Currently, there are no 'proteins' in DB. When accessing a gene, proteins sequences are …