-
The oyster genome has a gff annotation file with CGI_# as the identifier. The gtf file has a different set of LOC# and GeneID:#. Is there a file exists that links a CGI_# with one of the numbers in …
-
Hello
I'm trying to run LncPipe for the provided test data. Here is the command I run:
nextflow -c docker.config run ~/LncPipe/LncRNAanalysisPipe.nf --max_cpus 1 -with-trace -resume -with-timeli…
-
What I am seeing in the ioe file (generated from hg19 gtf file) is that Alternative First exons account for 42% of the splicing events. Skipped event exons follow at 23%. I was under the assumption t…
-
This is an attempt to have a common place to discuss on the unification task. Nothing is set in stone but at least let's try to converge on a common place to discuss.
What we have so far (not prete…
-
I run LncRNAanalysisPipe using your test data. It seems not to be completed.
```
$ nextflow run LncRNAanalysisPipe.nf -profile standard,docker,run
N E X T F L O W ~ version 20.01.0
Launching `L…
-
Hi to all!
When change SR in "settings options chane SR" apply we obtain that python error:
RuntimeError: wrapped C/C++ object of type ReseauTi has been deleted
Traceback (most recent call last):…
-
see https://github.com/r-transit/tidytransit/issues/106 for more
-
[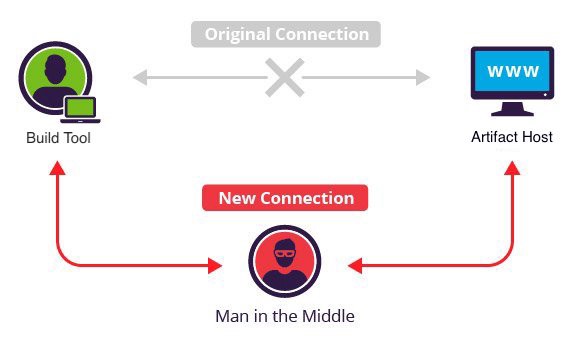](https://medium.com/@jonathan.leitschuh/want-to-take-over-the-java-ecosystem-all-yo…
-
Hi, I noticed that there was mention of an exon-based comparison tool but cannot find that option in the talon commands. I am trying to quantify differences in 3' and 5' exon boundaries compared with …
-
When enabling or disabling transit modes in an analysis, all routes added by scenarios are treated as trams.
There is no UI element to set the mode of a newly added route, but when an AddTrips obje…
abyrd updated
5 years ago