Cancer - Normal Transformer
Repo structure:
generate_figures.R :: Script generating the final figures for the paper
data/ :: Directory containing the data and upstream preprocessing scripts
analysis/ :: Directory containing the analyses performed as part of this study
misc/ :: Misc data, helper functions, and scriptsTLDR
TLDR: This repo is for a research paper that develops 2-3 approaches for addressing one question: "How do cancers relate to normal tissues?" In another sense, this question is asking:
- What are the normal cell states that a tissue can inhabit?
- What are the normal states which each cancer most resembles?
- How do interventions, such as chemotherapy, change this relationship?
- What new drug targets does this reveal? Does this tell us anything about the cell of origin?
Here are the three aims:
- Manifold Alignment (Aim 1): Generation of normal reference manifold and then alignment of "query" cancer samples to this reference. This allows us to visually see where in the reference they align and then compare them to eachother using techniques like clustering. This approach isn't as accurate, but it's an absolute requirement for our tool since people need to visually see the alignment to understand it. Pro: Very easy to visualize and interpret. Con: It's not that accurate as dimensionality reduction causes lots of information loss.
- Ontology-based normal-tissue scores (Aim 2): We use NMF to find pathway-based latent dimensions that we can use to map cancer samples to the cell ontology and derive normal tissue scores for each cancer sample. E.g., Cancer sample X is 50% T-Cell and 50% Stem Cell -- gives a more quantitative approach to mapping query cancers to normal tissue references.
- Cancer-reference projection (Aim 3): Creation of a cancer reference which we can generate scores for in normal tissues. This is the inverse of 2 where the normal is now the query and the cancer is the reference. There's certain reasons why this might actually be a more accurate strategy, but definitely still underdeveloped.
Background
This repository is based on a simple idea:
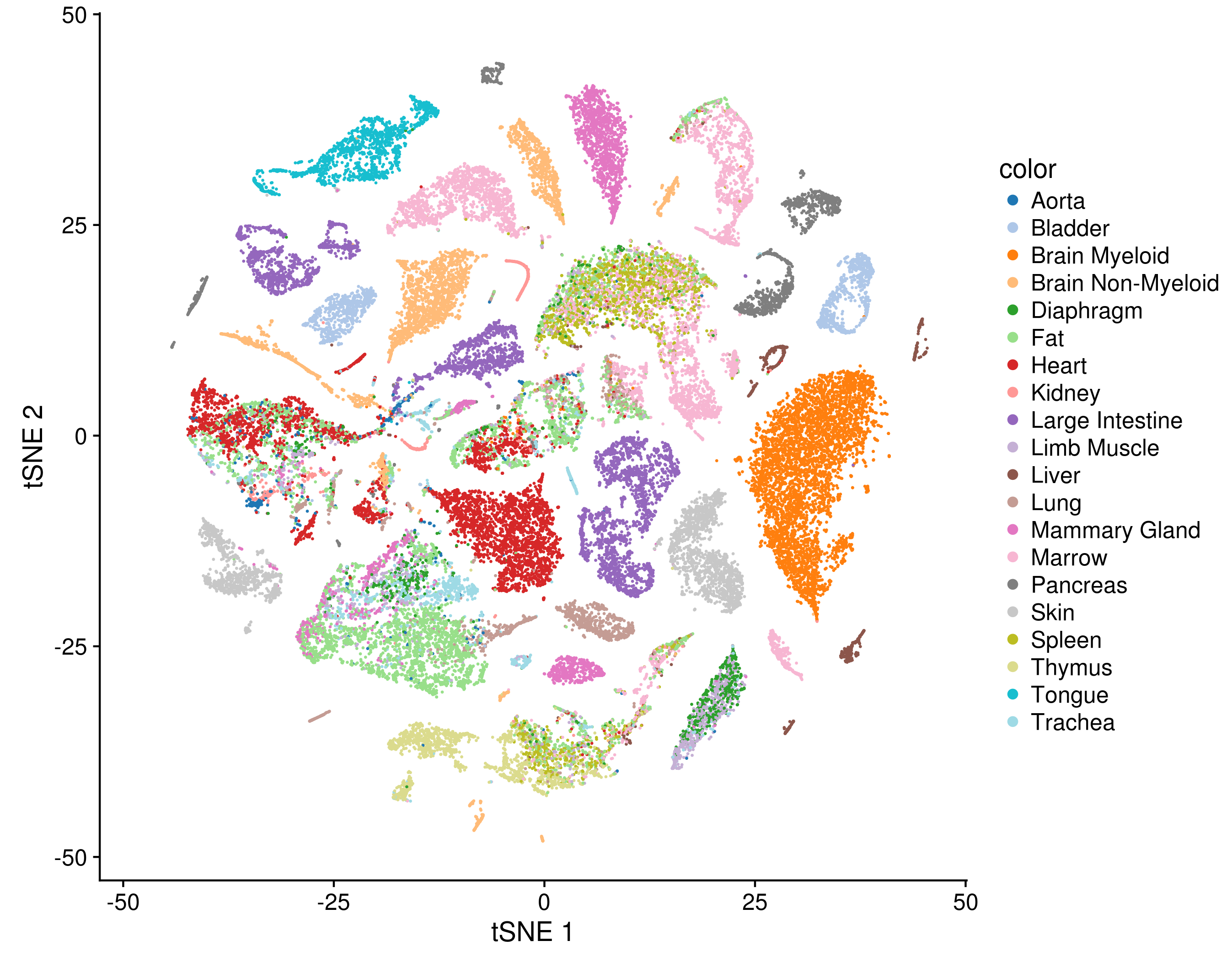
There is are a discrete number of Normal cellular states which are defined by the expression of specific genes in specific pathways -- and that we can learn these states using RNA-Sequencing. One popular depiction of this idea is the Tabula Muris - article:
From our recent Cancers paper, we showed that at least one cancer (Ewing sarcoma) has a cellular state which is similar to the cellular states found during normal embryonic development. More improtant, we learned how certain interventions change the state of Ewing sarcoma tumor cells. This led us to realize new drug targets which were subsequently demonstrated (albeit not by us...). However, the method we used (PHATE) is more qualitative with respect to answer questions of the normal-cancer relationship. For a larger-scale implementation of this approach, we need more quantitative tools in addition to qualitative ones.
The approach
A Cancer PHATE Atlas
We plan to extend the method we showed previously to all other cancers. Then, we hope to make a web application where users can explore their cancer of interest and see how it relates to normal tissues.
Cancers scored with normal tissue levels
Using a method similar to PLIER, we can incorporate pathway-level information to score cancer samples based on their expression of pathways which are related to particular normal tissues. For example, we might have found that Ewing sarcoma has a score of 50% Mesodermal and 50% pluripotency. How does an intervention change these scores? This would give us an innovative and quantitative way to assess the same questions posed above.
Normal tissues scores based on cancer signatures
This Dr. Zheng's idea which may provide an interesting supplement to our other analyses -- but he hasn't yet disclosed the details of his approach to me.