ndx-bipolar-scheme Extension for NWB
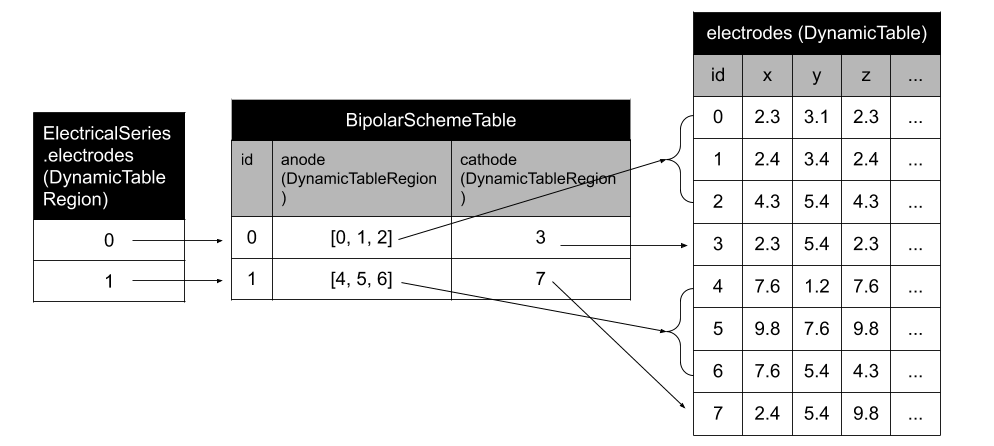
Structure for storing the bipolar schema of a recording in an NWB file.
python installation
$ pip install ndx-bipolar-schemepython usage
import os
from pynwb import NWBHDF5IO, NWBFile
from pynwb.file import DynamicTableRegion
from datetime import datetime
from ndx_bipolar_scheme import BipolarSchemeTable, NdxBipolarScheme
from pynwb.ecephys import ElectricalSeries
import numpy as np
nwbfile = NWBFile('description', 'id', datetime.now().astimezone())
device = nwbfile.create_device('device_name')
electrode_group = nwbfile.create_electrode_group('electrode_group',
'desc', 'loc', device=device)
for i in np.arange(20.):
nwbfile.add_electrode(i, i, i, np.nan, 'loc', 'filt', electrode_group)
electrodes = DynamicTableRegion(
name='electrodes',
data=np.arange(0, 3),
description='desc',
table=nwbfile.electrodes,
)
source_ec_series = ElectricalSeries(
name='source_ec_series',
description='desc',
data=np.random.rand(100, 3),
rate=1000.,
electrodes=electrodes,
)
nwbfile.add_acquisition(source_ec_series)
bipolar_scheme_table = BipolarSchemeTable(
name='bipolar_scheme', description='desc'
)
bipolar_scheme_table.add_row(anodes=[0], cathodes=[1])
bipolar_scheme_table.add_row(anodes=[0, 1], cathodes=[2, 3])
bipolar_scheme_table.add_row(anodes=[0, 1], cathodes=[2])
bipolar_scheme_table['anodes'].target.table = nwbfile.electrodes
bipolar_scheme_table['cathodes'].target.table = nwbfile.electrodes
bipolar_scheme_region = DynamicTableRegion(
name='electrodes',
data=np.arange(0, 3),
description='desc',
table=bipolar_scheme_table,
)
ec_series = ElectricalSeries(
name='dest_ec_series',
description='desc',
data=np.random.rand(100, 3),
rate=1000.,
electrodes=bipolar_scheme_region,
)
nwbfile.add_acquisition(ec_series)
ndx_bipolar_scheme = NdxBipolarScheme(
bipolar_scheme_tables=[bipolar_scheme_table],
source=source_ec_series
)
nwbfile.add_lab_meta_data(ndx_bipolar_scheme)
with NWBHDF5IO('test_nwb.nwb', 'w') as io:
io.write(nwbfile)
with NWBHDF5IO('test_nwb.nwb', 'r', load_namespaces=True) as io:
nwbfile = io.read()
nwbfile.acquisition['dest_ec_series'].electrodes.table['anodes'][2]['x']
os.remove('test_nwb.nwb')MATLAB usage
nwb = NwbFile( ...
'session_description', 'mouse in open exploration',...
'identifier', 'Mouse5_Day3', ...
'session_start_time', datetime(2018, 4, 25, 2, 30, 3), ...
'general_experimenter', 'My Name', ... % optional
'general_session_id', 'session_1234', ... % optional
'general_institution', 'University of My Institution', ... % optional
'general_related_publications', 'DOI:10.1016/j.neuron.2016.12.011'); % optional
nshanks = 4;
nchannels_per_shank = 3;
variables = {'x', 'y', 'z', 'imp', 'location', 'filtering', 'group', 'label'};
tbl = cell2table(cell(0, length(variables)), 'VariableNames', variables);
device = types.core.Device(...
'description', 'the best array', ...
'manufacturer', 'Probe Company 9000');
device_name = 'array';
nwb.general_devices.set(device_name, device);
device_link = types.untyped.SoftLink(['/general/devices/' device_name]);
for ishank = 1:nshanks
group_name = ['shank' num2str(ishank)];
nwb.general_extracellular_ephys.set(group_name, ...
types.core.ElectrodeGroup( ...
'description', ['electrode group for shank' num2str(ishank)], ...
'location', 'brain area', ...
'device', device_link));
group_object_view = types.untyped.ObjectView( ...
['/general/extracellular_ephys/' group_name]);
for ielec = 1:nchannels_per_shank
tbl = [tbl; {5.3, 1.5, 8.5, NaN, 'unknown', 'unknown', ...
group_object_view, [group_name 'elec' num2str(ielec)]}];
end
end
electrode_table = util.table2nwb(tbl, 'all electrodes');
nwb.general_extracellular_ephys_electrodes = electrode_table;
electrodes_object_view = types.untyped.ObjectView( ...
'/general/extracellular_ephys/electrodes');
electrode_table_region = types.hdmf_common.DynamicTableRegion( ...
'table', electrodes_object_view, ...
'description', 'all electrodes', ...
'data', (0:height(tbl)-1)');
source_electrical_series = types.core.ElectricalSeries( ...
'starting_time', 0.0, ... % seconds
'starting_time_rate', 30000., ... % Hz
'data', randn(12, 3000), ...
'electrodes', electrode_table_region, ...
'data_unit', 'volts');
nwb.acquisition.set('ElectricalSeries', source_electrical_series);
source_electrical_series_link = types.untyped.SoftLink( ...
'/acquisition/ElectricalSeries');
anodes_data = {0, [0, 1], [0, 1]};
cathodes_data = {1, [2, 3], 2};
[anodes, anodes_index] = util.create_indexed_column(anodes_data, ...
'/general/ndx_bipolar_scheme/bipolar_scheme', [], [], electrodes_object_view);
[cathodes, cathodes_index] = util.create_indexed_column(cathodes_data, ...
'/general/ndx_bipolar_scheme/bipolar_scheme', [], [], electrodes_object_view);
bipolar_scheme_table = types.ndx_bipolar_scheme.BipolarSchemeTable( ...
'id', types.hdmf_common.ElementIdentifiers('data', 0:2), ...
'description', 'my description', ...
'colnames', {'anodes', 'cathods'}, ...
'anodes', anodes, 'anodes_index', anodes_index, ...
'cathodes', cathodes, 'cathodes_index', cathodes_index);
ndx_bipolar_scheme = types.ndx_bipolar_scheme.NdxBipolarScheme(...
'bipolar_scheme', bipolar_scheme_table, ...
'source', source_electrical_series_link);
nwb.general.set('ndx_bipolar_scheme', ndx_bipolar_scheme);
nwbExport(nwb, 'test.nwb');