POSTRE
POSTRE: Prediction Of STRuctural variant Effects
LATEST UPDATE: 11/09/2024 What is new?
POSTRE can handle now Vision-Eye related phenotypes.
- What is POSTRE?
- How to use POSTRE?
- How to install and run POSTRE?
- Using POSTRE as an R function (alternative to usage with graphical user interface)
- Can we help you?
What is POSTRE?
Check the infographic displayed below to see a representation of POSTRE functionality. Click the image to expand it.
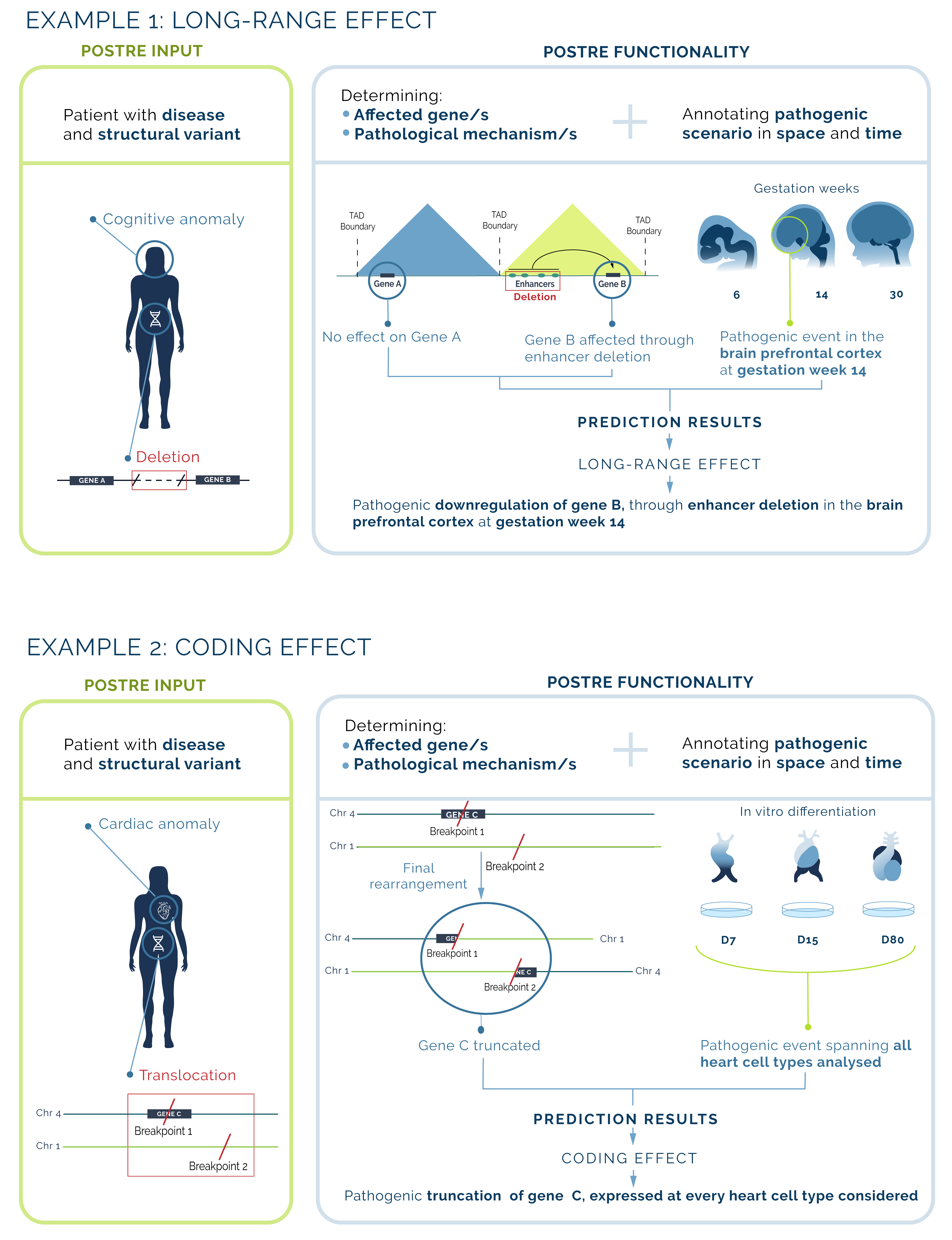
How to use POSTRE?
POSTRE can be used to analyse Single or Multiple SVs.
Analysing a Single SV
Watch POSTRE performance with a real patient in the following YouTube video. Reproduce it in Full Screen and High Quality (1080p) for optimal visualization. For this patient, with BOFS syndrome carrying an inversion (Laugsch et al., 2019), POSTRE successfully predicts the loss of TFAP2A expression in neural crest cells through an enhancer disconnection mechanism.
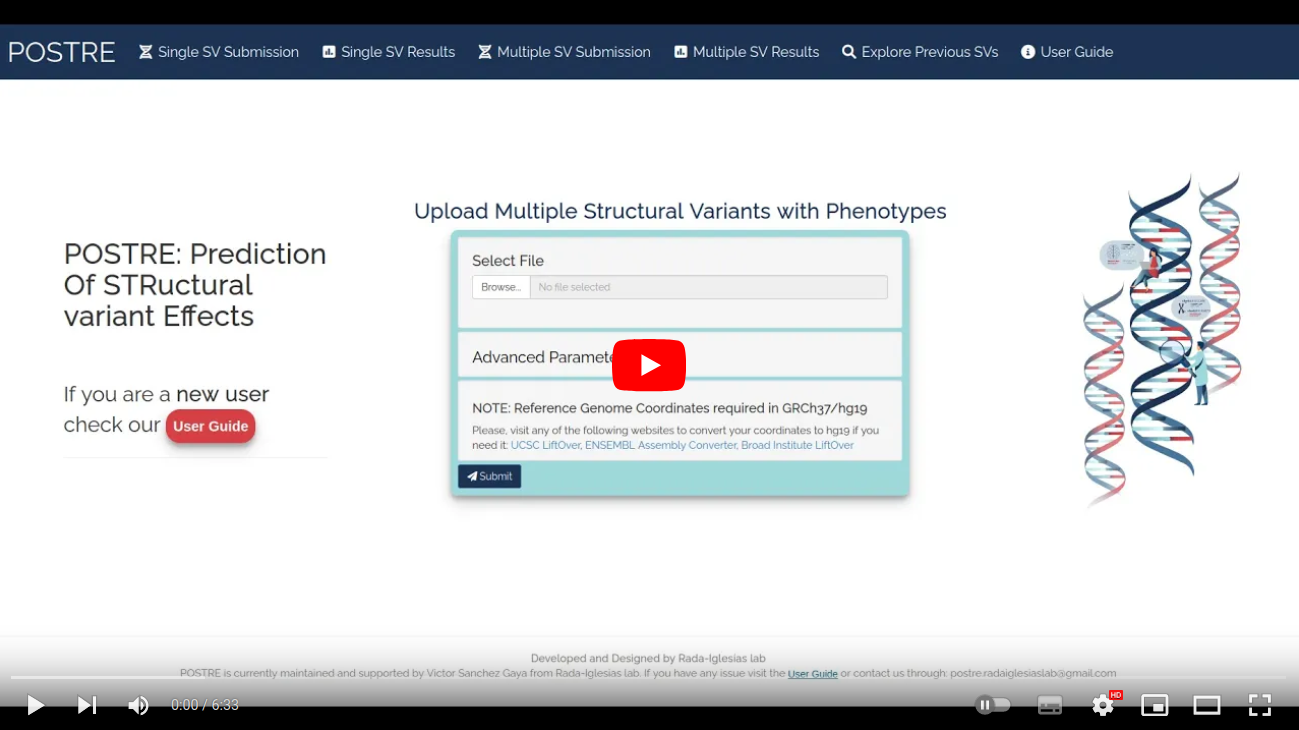
Analysing Multiple SVs
The Multiple SV Submission box allows the sequential analysis of multiple structural variants (results can be downloaded as txt tables). The SVs may come from just one or multiple patients. The SVs information has to be uploaded in a specific format. The file format consists on 1 line and 7 columns per structural variant. Each structural variant must contain only two breakpoints. The information associated with each column is provided below:
Note: For the case of structural variants happening strictly in one chromosome (deletions, inversions, duplications) the breakpoint 1 is the one associated with a smaller genomic coordinate, and the breakpoint 2 the one associated with a larger genomic coordinate. For translocations, it does not matter.
- Column 1: Chromosome for the breakpoint 1
- Column 2: Genomic coordinates (hg19) for the breakpoint 1. When not base pair resolution, provide a comma separated range, e.g. 85092268,85092269.
- Column 3: Chromosome for the breakpoint 2
- Column 4: Genomic coordinates (hg19) for the breakpoint 2. When not base pair resolution, provide a comma separated range, e.g. 85092268,85092269.
- Column 5: Structural Variant Type. Current options: Inversion, Translocation, Deletion or Duplication.
- Column 6: Comma separated list of phenotypes associated with the structural variant. Current options are: head_neck, limbs, neurodevelopmental, cardiovascular or vision_eye. For instance: head_neck,neurodevelopmental,cardiovascular.
- Column 7: Structural variant unique identifier e.g. (Patient1_SV3)
The data must be stored in a plain text file with column values separated by tabulations
An example file can be found here (to download it: (1) right-click the "Raw" button at the top of the file, (2) select Save Link As…, (3) choose the location on your computer where you want to save the file, and (4) select Save). Additional test files can also be downloaded from the testFiles folder.
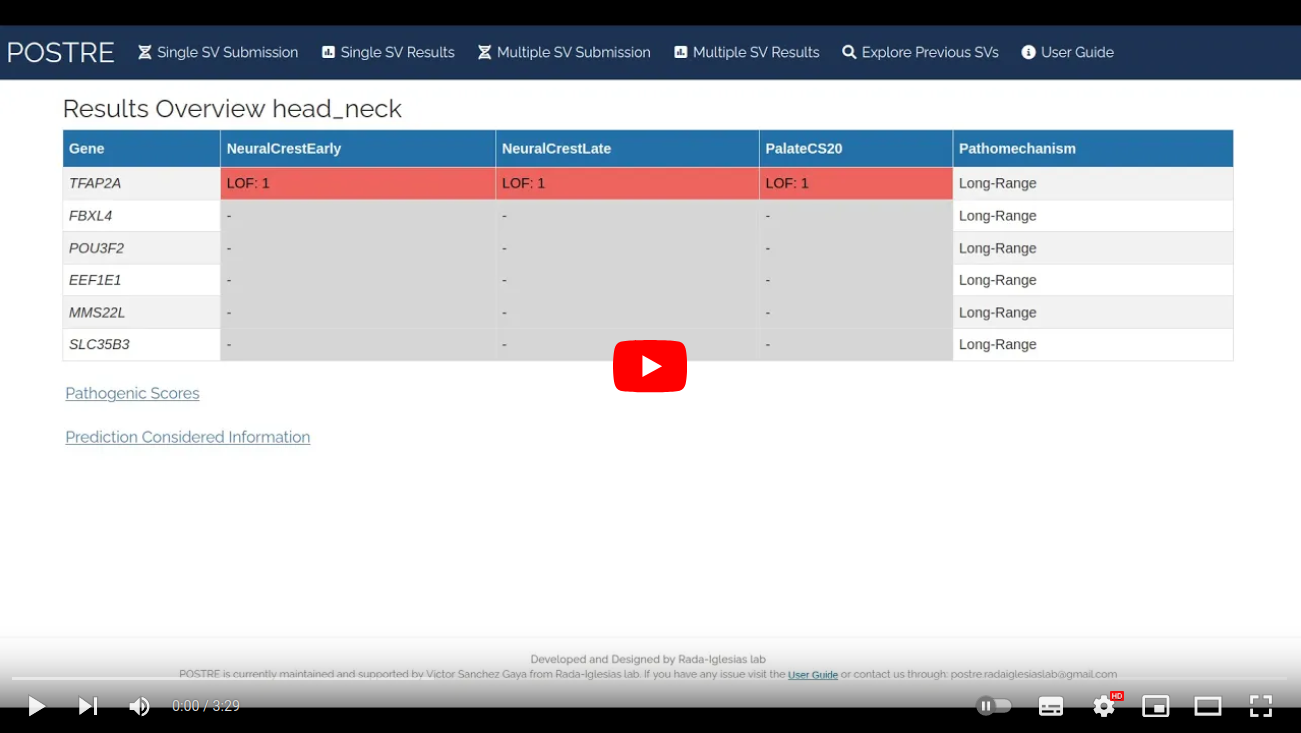
Upon the analysis of multiple SVs two main tables are provided. The first one (Results per SV and phenotype) is a table with a pathogenic prediction for each of the SVs and associated phenotypes analyzed. The second one (Results per gene and phenotype) is an aggregation of the pathogenic predictions per gene, phenotype and pathogenic mechanism (coding, long-range). Perform a Multiple SV Submission with one of the test files and check "How to navigate through this page?" section for more details.
A tutorial video showing how to perform and interpret results for a Multiple SV Submission analysis is provided in the following video.
How to install and run POSTRE?
Even though you may not have computational skills, do not be afraid!. POSTRE installation is very easy. On top of that, once POSTRE is installed, running and using it is as simple as any other desktop application. Thanks to its user friendly graphical interface.
POSTRE is built with Shiny framework. Thus, to run POSTRE you only require R (version >=3.5.0).
1. Installing R
To run Postre R version >=3.5.0 is required.
There is plenty of available information in the internet about how to install R depending on the OS (Windows, Mac, Linux etc.). Usually upon R installation people also install R-Studio, which is an integrated development environment to write R code. As a result, most tutorials explain how to install both R and R-Studio. But, to run Postre R-Studio is not necessary. If you need help for R download, different tutorials are provided here:
- Windows Youtube Tutorial
- Mac Youtube Tutorial
- Ubuntu Youtube Tutorial
- Web Tutorial for installing R and R-Studio in Windows, Mac or Linux
2. Running POSTRE Latest Version
Once R is installed, just execute the following instruction in an R console to run POSTRE latest version:
source("https://raw.githubusercontent.com/vicsanga/Postre/main/Postre_wrapper.R")The above instruction installs and loads all the required libraries for POSTRE. If some libraries are missing R may ask for permission to install them, confirm it. The first time that you run POSTRE this action will probably take more time.
If you are not sure about how to do it. You can find me initializing POSTRE from R in the video below!
Alternative option, POSTRE online
POSTRE can also be uploaded to a server (e.g. shinyapps.io), by a user with R installed, and accessed online as a normal web app by other users.
A detailed explanation on how to do that is provided in Shiny web page.
Recapitulating, you have to:
- Install the rsconnect R package
- Create an account at shinyapps.io
- Use the tokens generated by shinyapps.io to configure your rsconnect package.
- Deploy apps with rsconnect::deployApp. Regarding this step, download and uncompress POSTRE Github repository. POSTRE required information for the upload to shinyapps.io is located in Postre_app folder. Thus, this is the folder that has to bee synchronized with rsconnect. e.g: ```R rsconnect::deployApp('/pathInYourDeviceTo/Postre/Postre_app/', account = 'nameOfYourAccount') ```
The advantage of this approach is to provide other users in your organization access to the tool without having R installed. However, the person in charge of uploading POSTRE to the server will have to check for new versions or releases of the tool, and re-upload them to the server when available. Contrarily, if POSTRE is executed from an R terminal, with the source() command given above, in step 2, the latest version of POSTRE will always be executed. If you want to be notified when new major releases of the tool are available, please, send a mail to postre.radaiglesiaslab@gmail.com to be added to the notifications mail list.
Using POSTRE as an R function
If you want to use POSTRE as an R function, and not through its graphical user interface, you can do that by following the steps provided below. Importantly, if you want to take profit of POSTRE latest version, to update POSTRE R function you will have to repeat the steps 1 and 2 each time a new version is released. Latest version release date: 11/09/2024. If you want to be notified when new major releases of the tool are available, please, send a mail to postre.radaiglesiaslab@gmail.com to be added to the notifications mail list.
Steps for using POSTRE as an R function:
- Download POSTRE repository. For instance, compressed as .zip through the: "Code" GitHub button (green button at the top of this page) and uncompress it after downloading.
- In the R script where you want to use POSTRE R function, to load it, do a source of the POSTRE_multiSV.R script (i.e. source("/PathTo/Postre_app/POSTRE_multiSV.R")). This script is located inside of the "Postre_app" folder (script + folder are downloaded in Step 1). Example: ```R ##This is an R script ##Loading POSTRE R function source(file = "/home/victor/Downloads/Postre-main/Postre_app/POSTRE_multiSV.R") ```
- POSTRE R function, named POSTRE_multiSV(), is already loaded in the R environment. To use it, it requires 2 mandatory arguments.
- SVs: Data frame with 7 columns containing the SVs information (coordinates have to be in hg19). You can use as an example, to generate the data frame, any of the test files provided in the GitHub testFiles folder (you can find it at the top of this page). More information about how to define the SVs information is given in the Analysing Multiple SVs section.
- pathTo_Postre_app_Folder: Provide the path to "Postre_app" folder (downloaded in Step 1): i.e. pathTo_Postre_app_Folder="/home/victor/Downloads/Postre-main/Postre_app/"
Regarding the output, a list with three different elements is provided.- pathogenicityPrediction_per_SV: This table provides a pathogenic prediction for each of the SVs and associated phenotypes analyzed.
- geneStats_recurrencyAndPathomech: This table contains an aggregation of the pathogenic predictions per gene, phenotype and pathogenic mechanism (coding, long-range).
- errors: Contains the ids of the SVs which rised an error (if they occur) during their interpretation.
If you want to know more details about the different parameters (mandatory and optional), and output provided by the function, run the function without arguments: ```R ################################################### ##Getting all details about parameters and output POSTRE_multiSV() ``` Here is provided a full example of how to use the function with default parameters: ```R ############################## ## Example of function Usage ############################## ##Loading POSTRE R function (named: POSTRE_multiSV()) source(file = "/home/victor/Downloads/Postre-main/Postre_app/POSTRE_multiSV.R") #Loading SVs patients svs_patients<-read.delim(file = "/home/victor/Downloads/Postre-main/testFiles/Table1_LongRange_SVs.tsv", sep="\t", stringsAsFactors = FALSE, header = FALSE) #Running POSTRE R function, with default parameters res_svs<-POSTRE_multiSV(SVs = svs_patients, pathTo_Postre_app_Folder = "/home/victor/Downloads/Postre-main/Postre_app/") #Visualizing data frame with pathogenic prediction per SV and associated phenotype View(res_svs$pathogenicityPrediction_per_SV) ```
Can we help you?
If you are experiencing some trouble, or you just want to contact us for any reason, please send a mail to postre.radaiglesiaslab@gmail.com.