molplotly
molplotly is an add-on to plotly built on RDKit which allows 2D images of molecules to be shown in plotly figures when hovering over the data points.
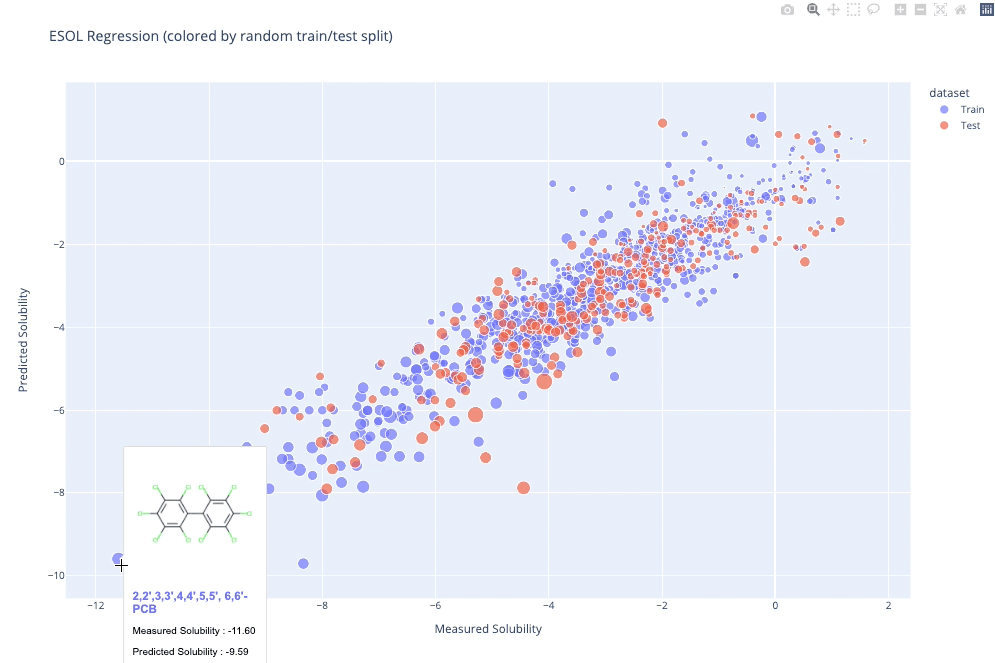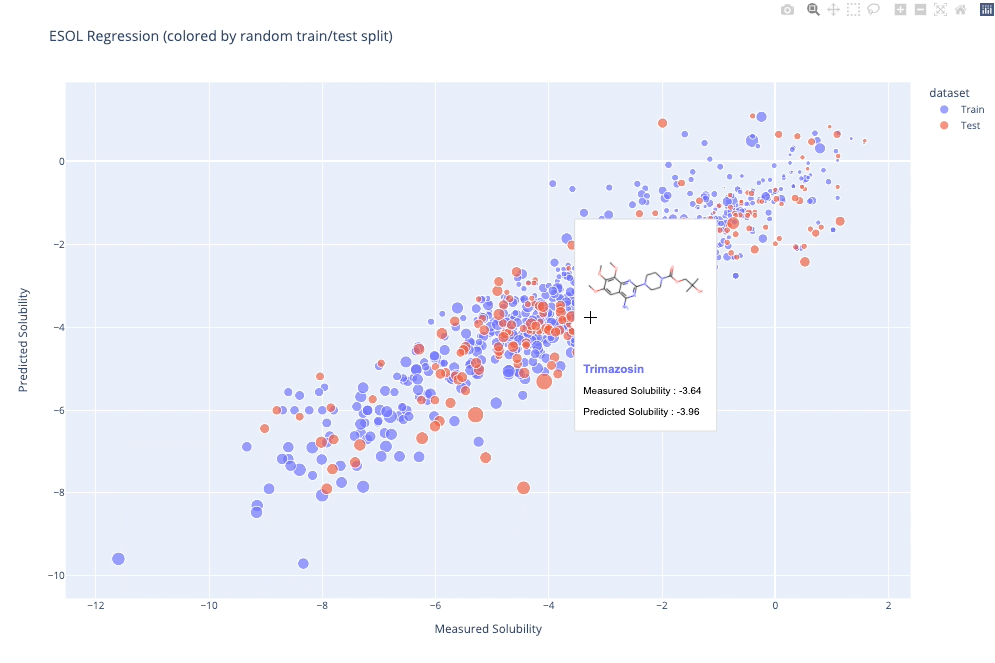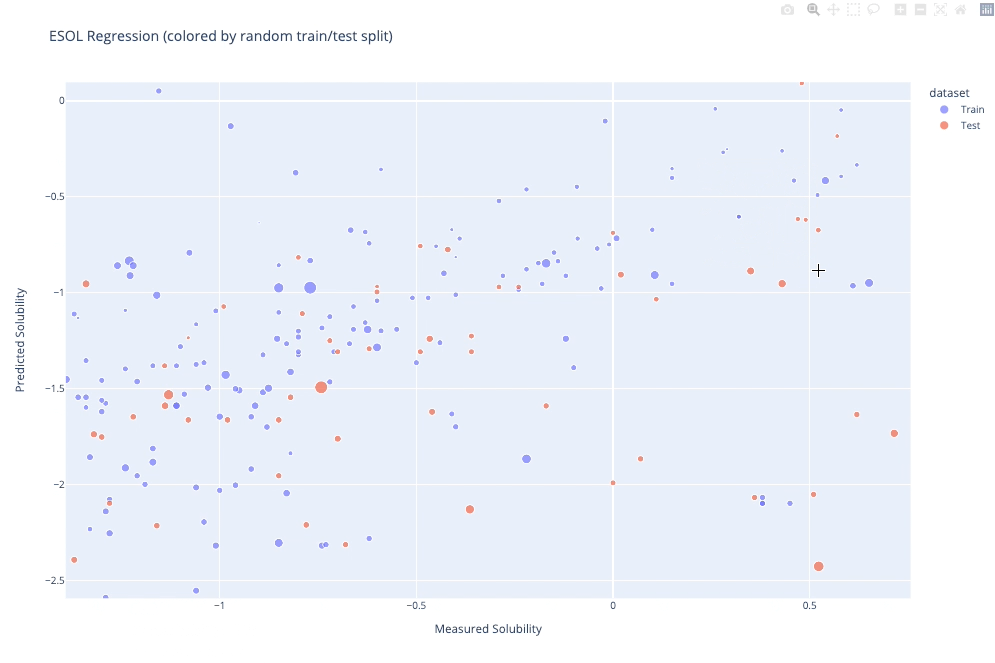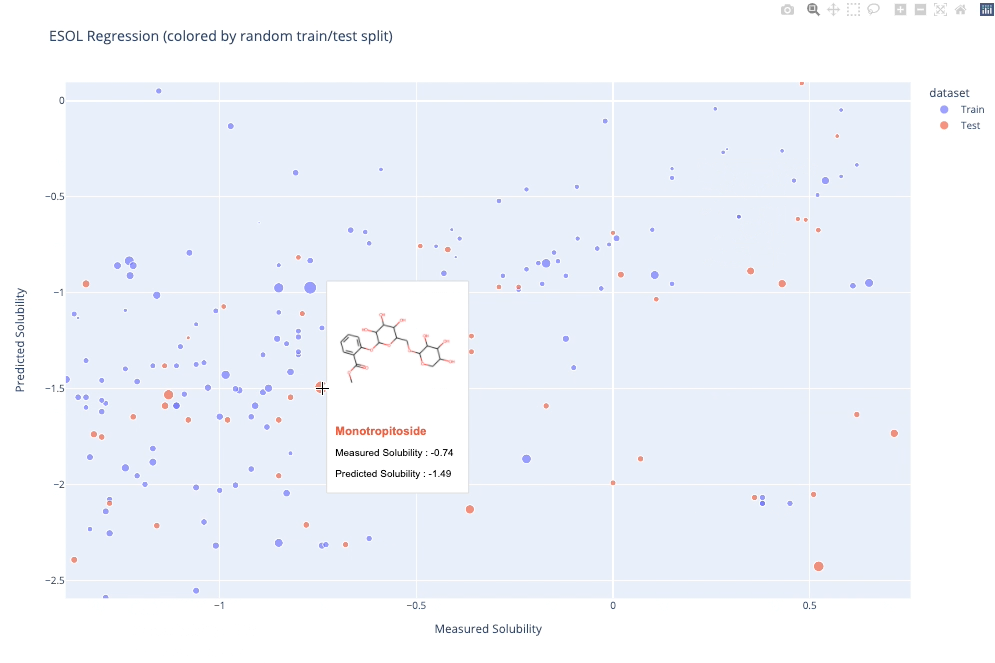
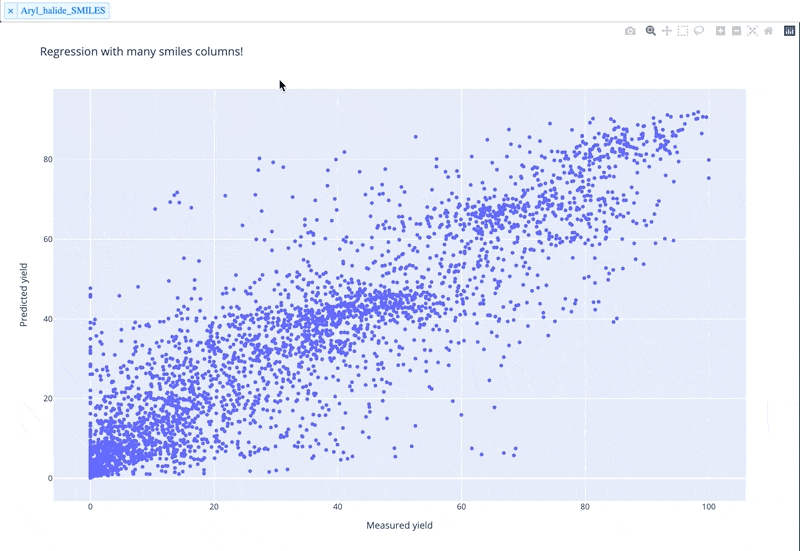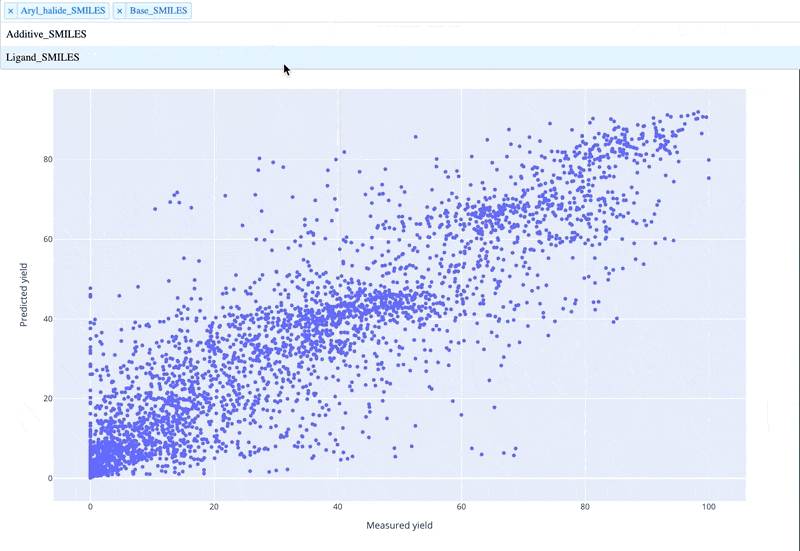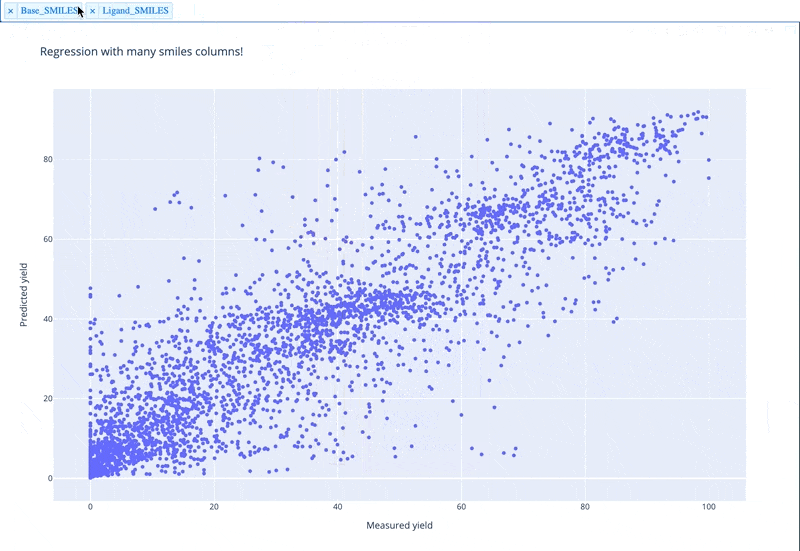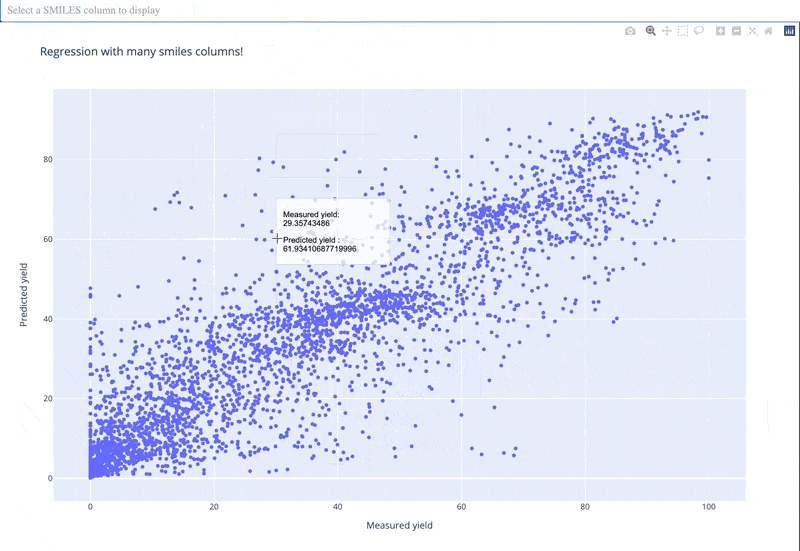
A readable walkthrough of how to use the package together with some useful examples can be found in this blog post while a runnable notebook can be found in examples/simple_usage_and_formatting.ipynb :)
Installation
pip install molplotlyUsage
import pandas as pd
import plotly.express as px
import molplotly
# load a DataFrame with smiles
df_esol = pd.read_csv(
'https://raw.githubusercontent.com/deepchem/deepchem/master/datasets/delaney-processed.csv')
df_esol['y_pred'] = df_esol['ESOL predicted log solubility in mols per litre']
df_esol['y_true'] = df_esol['measured log solubility in mols per litre']
# generate a scatter plot
fig = px.scatter(df_esol, x="y_true", y="y_pred")
# add molecules to the plotly graph - returns a Dash app
app = molplotly.add_molecules(fig=fig,
df=df_esol,
smiles_col='smiles',
title_col='Compound ID',
)
# run Dash app inline in notebook (or in an external server)
app.run_server(mode='inline', port=8700, height=1000)Input parameters
| name | type | default | description |
|---|---|---|---|
fig |
figure |
required | a plotly figure object containing datapoints plotted from df. |
df |
DataFrame |
required | a pandas dataframe that contains the data plotted in fig. |
smiles_col |
str |
'SMILES' |
name of the column in df containing the smiles plotted in fig |
show_img |
bool |
True |
whether or not to generate the molecule image in the dash app |
svg_size |
float |
200 |
the size in pixels of the molecule drawing |
alpha |
float |
0.7 |
the transparency of the hoverbox, 0 for full transparency 1 for full opaqueness |
mol_alpha |
float |
0.7 |
the transparency of the SVG molecule image, 0 for full transparency 1 for full opaqueness |
title_col |
str |
None |
name of the column in df to be used as the title entry in the hover box |
show_coords |
bool |
True |
whether or not to show the coordinates of the data point in the hover box |
caption_cols |
list |
None |
list of column names in df to be included in the hover box |
caption_transform |
dict |
{} |
Functions applied to captions for formatting. The dict must follow a key: function structure where the key must correspond to one of the columns in subset or tooltip |
color_col |
str |
None |
name of the column in df that is used to color the datapoints in df - necessary when there is discrete conditional coloring |
symbol_col |
str |
None |
name of the column in df that is used to determine the marker shape of the datapoints in df |
wrap |
bool |
True |
whether or not to wrap the title text to multiple lines if the length of the text is too long |
wraplen |
int |
20 |
the threshold length of the title text before wrapping begins - adjust when changing the width of the hover box |
width |
int |
150 |
the width in pixels of the hover box |
fontfamily |
str |
'Arial' |
the font family used in the hover box |
fontsize |
int |
12 |
the font size used in the hover box - the font of the title line is fontsize+2 |
Output parameters
by default a JupyterDash app is returned which can be run inline in a jupyter notebook or deployed on a server via app.run_server()
- The recommended
heightof the app is50+(height of the plotly figure). - For the
portof the app, make sure you don't pick the sameportas anothermolplotlyplot otherwise the tooltips will clash with each other. Also, apparently on windows port numbers below8700are used by other processes so for safety processes keep to numbers above that.
Can I run this in colab?
JupyterDash is supposed to have support for Google Colab but at some point that seems to have broken.. Keep an eye on the raised issue here! Update (1st March 2022): The plots seem to be running again but the hoverboxes are not showing so I don't think it has been fully fixed - I will keep an eye on it in the meantime.
Can I save these plots?
An issue/feature request for this has already been raised here.
moltplotly works using a Dash app which is non-trivial to export because server side javascript is needed in addition to HTML/CSS styling (as detailed here)
Until I find a way to get around that, the best alternative is to either host the plot on an app/server, exporting the plotly figure without molecules showing :( as detailed in this page. If you want to use it in a presentation I'd suggest keeping the figure open in a browser and changing windows to it during your talk!
Warning about memory size
Just adding a warning here that memory usage in a notebook can increase significanly when using plotly (not molplotly's fault!). If you notice your jupyter notebook slowing down, plotly itself is a likely culprit... In that case I'd consider either using plotly with static image rendering, or ... use seaborn :P

