-
Hi,
I'm having a problem with ciftify_recon_all and then ciftify_subject_fmri as well. I ran recon-all in FS version 7.1.0 and used the outputs of that for ciftify_recon_all, and I get this error:…
-
Dear alegiac95,
thanks for providing the scripts! I have just gone through the paper and description of this GitHub repo and I want to adapt your software to my project. However, I use the typical …
-
I have started testing the runtime for a more practical number of total streamlines/seeds. I used 10 million seeds (`-seeds 10M`) which in turn resulted in ~2.2M streamlines remaining after ACT.
He…
-
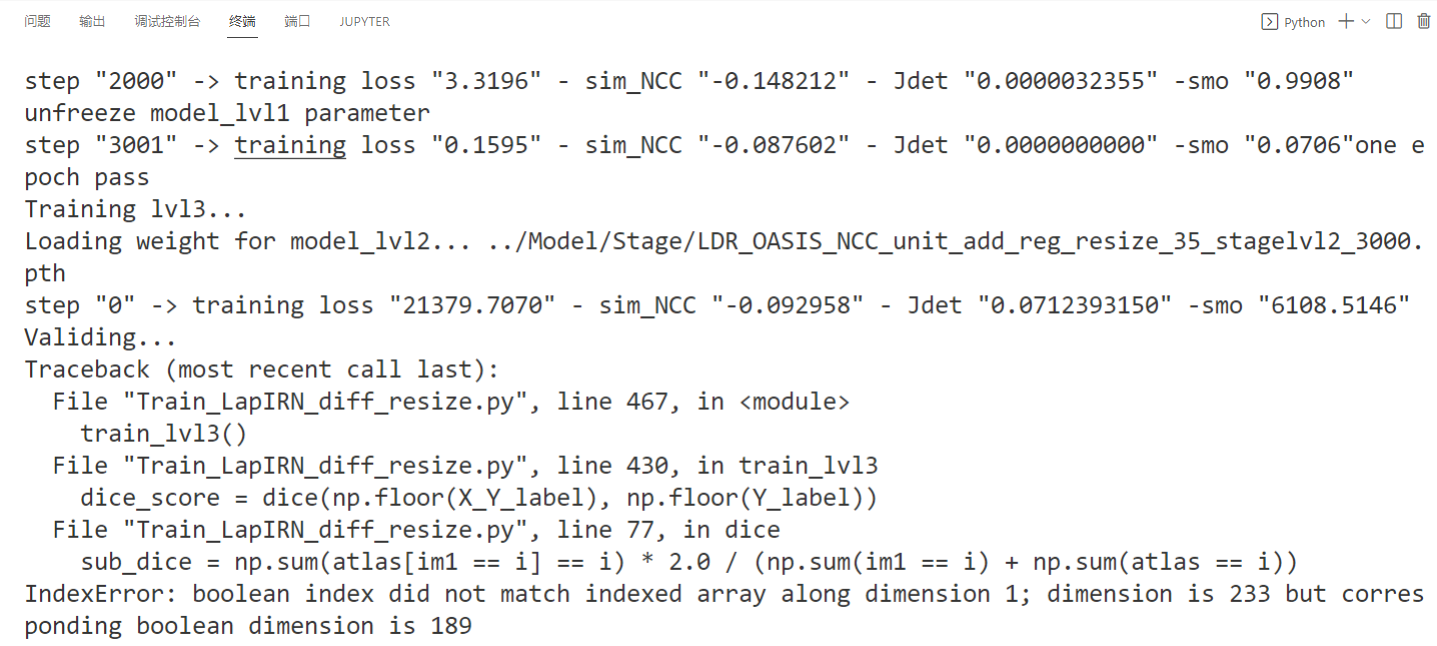
I'm sorry to bother you. When I used my data set to reproduce your code, there was such…
-
Related to https://github.com/mne-tools/mne-python/issues/10090.
In working with my own data, we have contacts that are different sizes and using a strict distance-based metric of labeling as imple…
-
The original atlases (both cortical and subcortical) are stored without any intensity scaling (multiplier of 1):
```
$ mrinfo /home/sina/Documents/Research/Datasets/UK_biobank/sample/1000243_2_0/d…
-
## Resource info table
| | |
| :------------------ | :------------------------…
-
If I try to load the Harvard Oxford sub-maxprob-thr0-1mm atlas with the following code:
```
from nilearn import input_data
from nilearn import datasets
atlas = datasets.fetch_atlas_harvard_oxf…
-
### Describe the bug
Build_nuisance_regressor crash: number of time points is inconsistent
```Node: cpac_103430-100_3_ses-1.nuisance_regressors_default_271.build_nuisance_regressors
Working direc…
-
Hello!
I'm coregistering patients' T1 to template. The two patients I analyzed both had subcortical lesion, but the lesion mask contained cortical patches. I wonder if there is a way to correct f…