PAMNet: A Universal Framework for Accurate and Efficient Geometric Deep Learning of Molecular Systems
Official implementation of PAMNet (Physics-aware Multiplex Graph Neural Network) in our paper A universal framework for accurate and efficient geometric deep learning of molecular systems accepted by Nature Scientific Reports.
PAMNet is an improved version of our MXMNet and outperforms state-of-the-art baselines regarding both accuracy and efficiency in diverse tasks including small molecule property prediction, RNA 3D structure prediction, and protein-ligand binding affinity prediction.
This implementation is also applicable to:
- Efficient and Accurate Physics-aware Multiplex Graph Neural Networks for 3D Small Molecules and Macromolecule Complexes (preprint).
- Physics-aware Graph Neural Network for Accurate RNA 3D Structure Prediction (Machine Learning for Structural Biology Workshop at NeurIPS 2022).
If you have any questions, feel free to open an issue or reach out to: szhang4@gradcenter.cuny.edu.
Overall Architecture
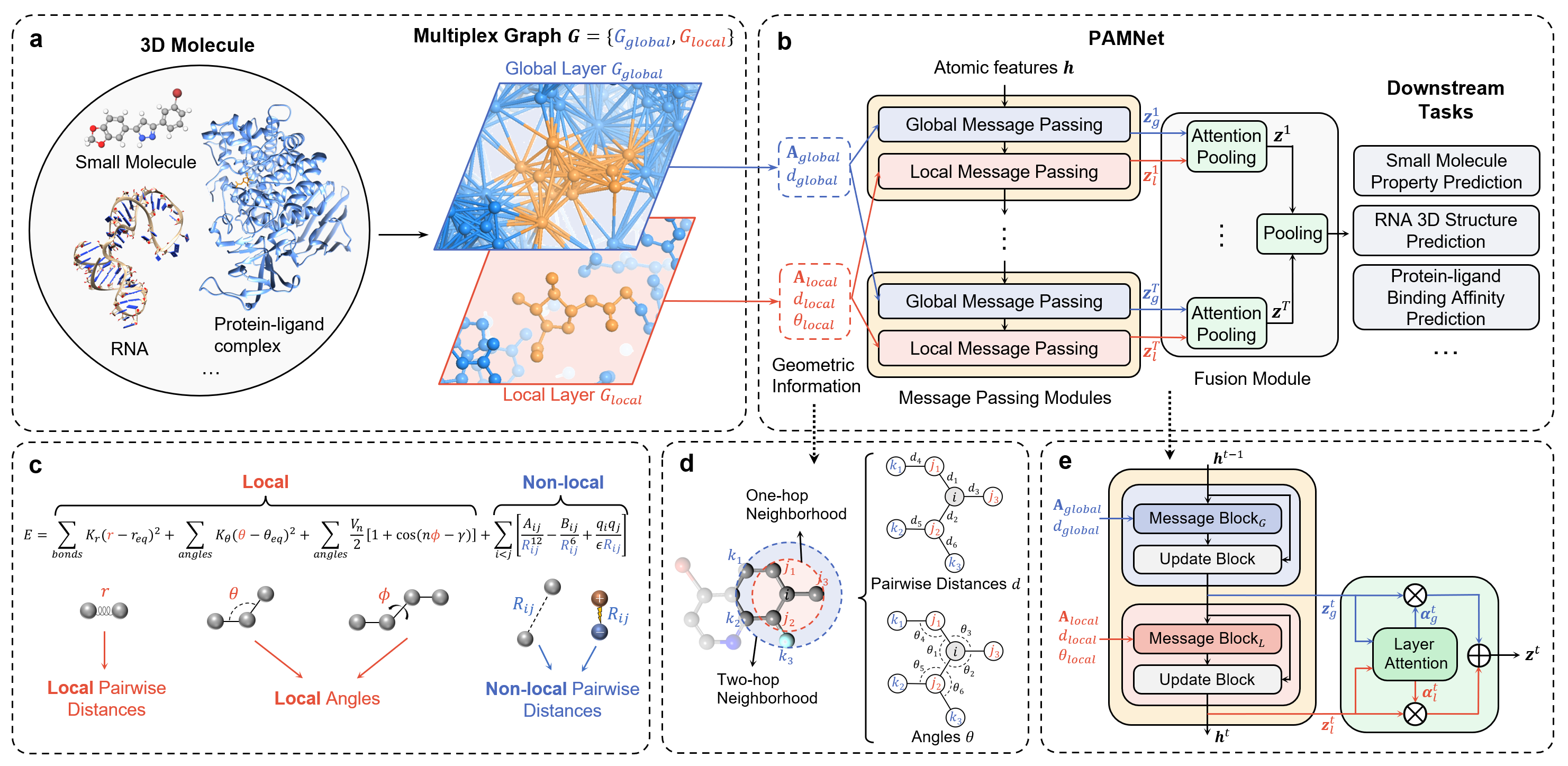
Updates
2024-01We provide the docker image for running PAMNet at https://hub.docker.com/r/zetayue/pamnet.2023-10PAMNet paper was accepted by Nature Scientific Reports.2023-07We release the code for PAMNet.
Environment Setup
Option 1: Base on Dependencies
- Python : 3.7.4
- CUDA : 10.1
Dependencies can be installed with:
pip install -r requirements.txtOptional: Install Open Babel 3.1.1 for binding affinity prediction on PDBbind:
- Download source file
conda install filename
Option 2: Docker Image
Docker image for running PAMNet is available at https://hub.docker.com/r/zetayue/pamnet.
Requires NVIDIA Container Toolkit installed to run with GPU support.
Example command to run:
nvidia-docker run --name pamnet -it --network=host --shm-size=1g --rm -v LOCAL_PATH:MOUNTED_PATH zetayue/pamnet:latestDatasets
QM9 for small molecule property prediction:
The training script (main_qm9.py) will automatically download the QM9 dataset and preprocess it.
PDBbind for protein-ligand binding affinity prediction:
- Download
PDBbind_dataset.tar.gzfrom dropbox - Unzip the downloaded file under
./data/PDBbind. There will be two subfolders (core-setandrefined-set) after the unzip - Run
python preprocess_pdbbind.pyto preprocess the dataset to construct graphs
RNA-Puzzles for RNA 3D structure prediction:
- Download
classics_train_val.tarfrom Stanford Digital Repository - Unzip the downloaded file under
./data/RNA-Puzzles. There will be one subfolderclassics_train_valcontainingexample_trainandexample_valafter the unzip - Run
python preprocess_rna_puzzles.pyto preprocess the dataset to construct graphs
How to Run
Arguments
--gpu GPU number
--seed random seed
--dataset dataset to be used
--epochs number of epochs to train
--lr initial learning rate
--wd weight decay value
--n_layer number of hidden layers
--dim size of input hidden units
--batch_size batch size
--cutoff_l distance cutoff used in the local layer
--cutoff_g distance cutoff used in the global layer
--model model to be used on QM9
--target index of target (0~11) for prediction on QM9Example command for training and evaluation
Small molecule property prediction on QM9:
python -u main_qm9.py --dataset 'QM9' --model 'PAMNet' --target=7 --epochs=900 --batch_size=32 --dim=128 --n_layer=6 --lr=1e-4Protein-ligand binding affinity prediction on PDBbind:
python -u main_pdbbind.py --dataset 'PDBbind' --epochs=170 --batch_size=32 --dim=128 --n_layer=3 --lr=1e-3RNA 3D structure prediction on RNA-Puzzles:
python -u main_rna_puzzles.py --dataset 'RNA-Puzzles' --epochs=15 --batch_size=8 --dim=16 --n_layer=1 --lr=1e-4Citation
If you find our model and code helpful in your work, please consider citing us:
@article{zhang2023universal,
title={A Universal Framework for Accurate and Efficient Geometric Deep Learning of Molecular Systems},
author={Zhang, Shuo and Liu, Yang and Xie, Lei},
journal={Scientific Reports},
volume={13},
number={1},
pages={19171},
year={2023},
publisher={Nature Publishing Group UK London}
}
@article{zhang2022physics,
title={Physics-aware graph neural network for accurate RNA 3D structure prediction},
author={Zhang, Shuo and Liu, Yang and Xie, Lei},
journal={arXiv preprint arXiv:2210.16392},
year={2022}
}
