-
There is a section in "tools" in which a `compute_pairwise_coalescence_times.py` script is used to calculate pairwise coalescent rates between one haplotype and the others. But there is no file called…
-
There is a bug in the simul_coalescent_only. :-1: And perhaps in spatialCoalescentSimulation too :-1: :-1:
Sometimes, it seems that several nodes can arrive in a deme with 0 carrying capacity. He…
-
In contrast to many methods, the combination of tsinfer + tsdate-vgamma does not incorporate a coalescent prior (it has a improper "flat" prior on most nodes and an exponential prior on the roots, fit…
-
* Skygrid
* Comparison of:
* Piecewise constant
* Continuous and piecewise linear
* Novel smooth population function?
-
A population can have an activation time that is greater than zero -- for example, to sample ancient individuals. When using the demography debugger's `coalescence_rate_trajectory()` function, specif…
-
library(mlesky)
treeMRSA
-
Would it be possible to have SINGER output the "full ARG" with marked recombination nodes (similar to that from msprime)? This would require maintaining unary nodes in the tree sequence, such as recom…
-
**Describe the bug**
A clear and concise description of what the bug is.
the program fails to compile
**To Reproduce**
Give exact command you ran. When you run ASTRAL, make sure you keep the s…
-
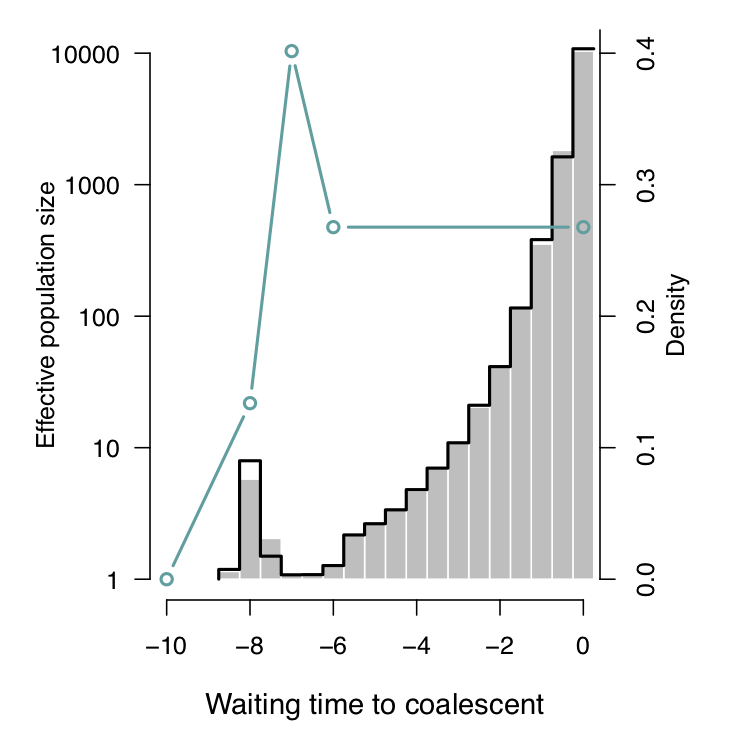
Slight but reproducible discordance around time point -8, why?
-
This would make a nice basis for #42