-
It seems that "dendogram + heatmap" are already possible to use in plotlyjs:
https://plot.ly/~talgalili/23
Related PR: https://github.com/plotly/plotly.js/pull/1063
Unanswered question on "commun…
-
https://github.com/reconstrue/neuro_on_colab/issues/19
- Clustergrammer do that? See #34
-
Can you create a dendrogram from the dist results?
Also, could you recommend parameters for large fungal genome comparison?
-
How would we interpret the dendogram output? If you can explain with an example
-
Version (Anaconda): drep 3.5.0 pyhdfd78af_0 bioconda
First of all, thanks for your work, dRep has been very useful in my work.
One thing that could be improv…
-
-
Would it be possible to remove margin (on the left side of the plot), because it makes the plots in the online docs function badly.
When you open the plot in the browser you can see the dendogram p…
-
users should be able to view a set of annotated entities (genes, genotypes, diseases, whatever) in a dendogram viewer.
http://bl.ocks.org/mbostock/4063570
this could be separate or in addition to th…
-
https://piziadas.com/en/hugo
>HUGO (Highly Uniform Graphic Organization) is a software tool; it has been developed to generate graphical representations of neurons stored in Neuromorpho. HUGO is a JA…
-
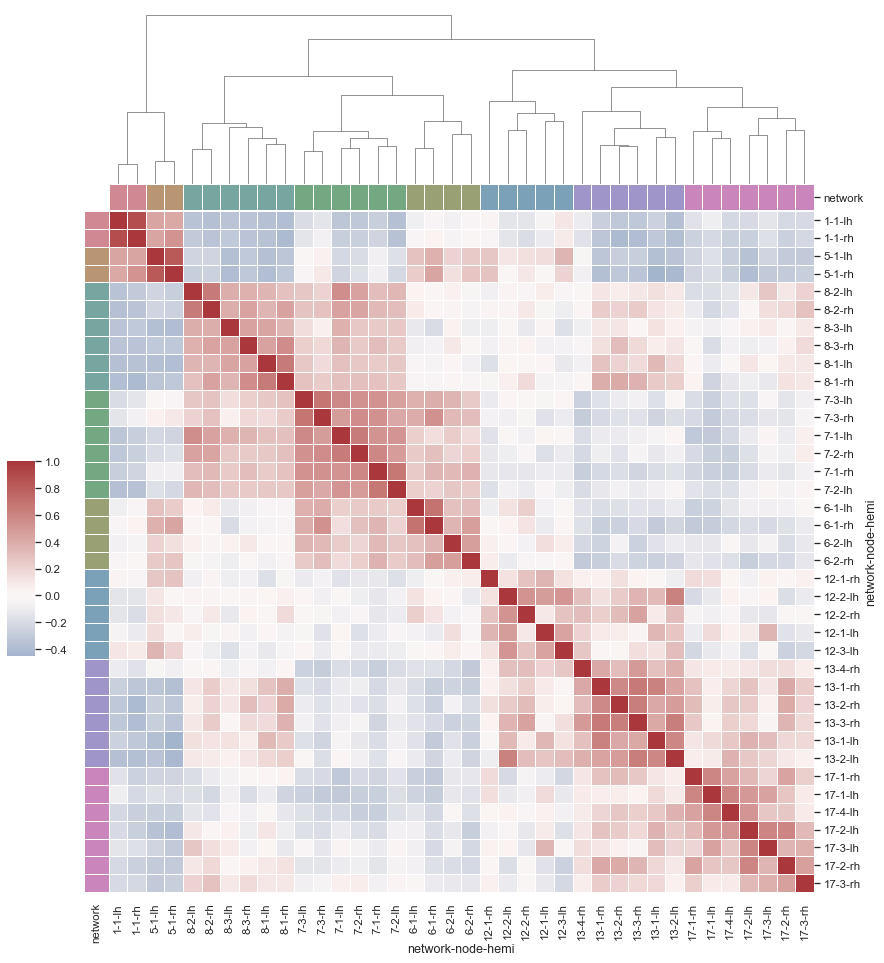
Hello! I would like to create a similar plot to the one shown below from [Seaborn's documentation](https://seaborn.pydata.org/examples/structured_heatmap.html). I understand that there are functions …